Adding colab notebook demo and an interface#3
Adding colab notebook demo and an interface#3idhamari wants to merge 18 commits intocwmok:masterfrom
Conversation
|
Dear @cwmok thanks for sharing. I noticed when running the notebook that the training loss is a large negative value e.g. Is this normal? |
|
Hi idhamari, This is not normal. I think the problem could be related to your dataset/preprocessing pipeline. Did you normalize the intensity of your data, i.e.: [0, 1]? Please also check out the related issue at #2 . |
|
Hi @cwmok thanks for your quick reply. I am using your code without modification and one of the original challenge public datasets. I think there is already normalization process in the generator class |
|
Thanks for spotting it out. The zero-mean 1-std normalization is not working with the normalized cross-correlation function (the background intensity of the image should be 0). During my experiment, I preprocess the training data within [0,1] and turn the "norm=False". I will update the code very soon. I apologize for the confusion. |
|
@cwmok thanks again for the clarification. I modified the code as you suggested and also added a normalisation code. However, I still get a negative loss value (it is smaller now). I also get zero Jacobian determinant. e.g. Maybe there is still there is something missing? |
|
Hi @idhamari, I found that the "normalization code" is not executed as I set |
|
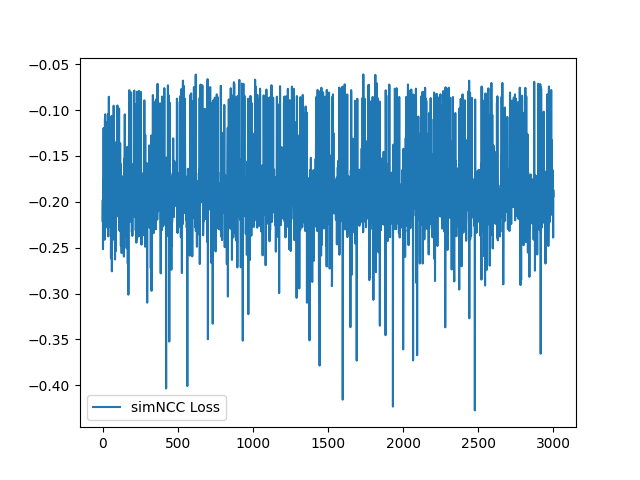
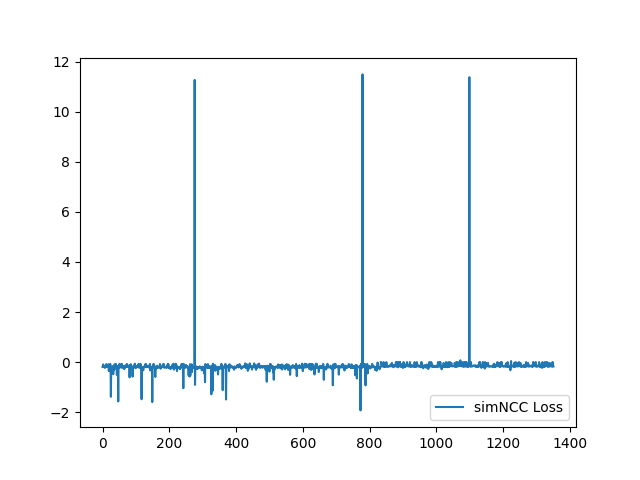
Now the loss function looks good. In our paper, we use negative NCC and therefore the value of sim_NCC range from -1 to 0. |
|
@cwmok Thanks for your comment.
Yes, it is clear now.
I am confused, isn't this conflict with:
more questions :)
so it is normal that Jdet = 0 ? I also noticed that the training loss is going up down, is this also expected? |
|
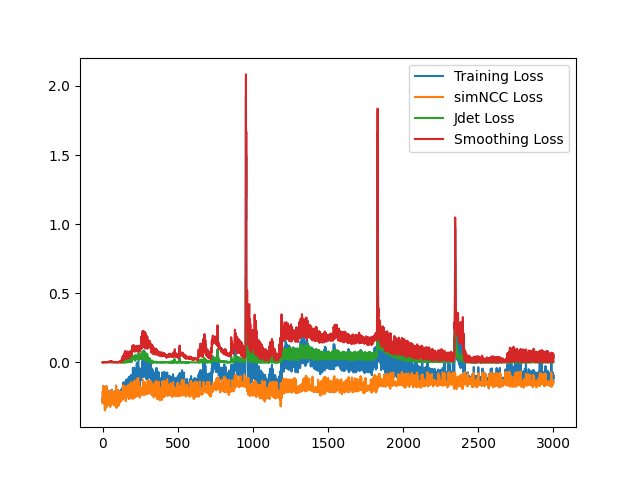
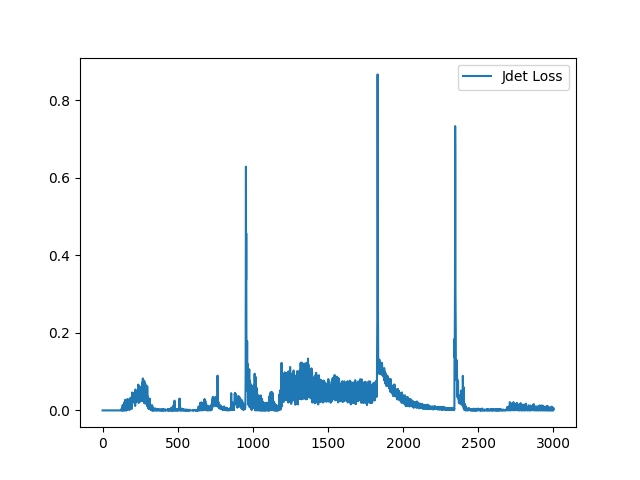
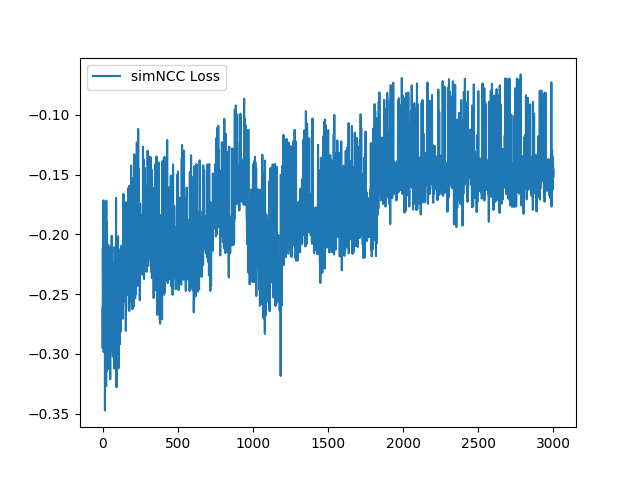
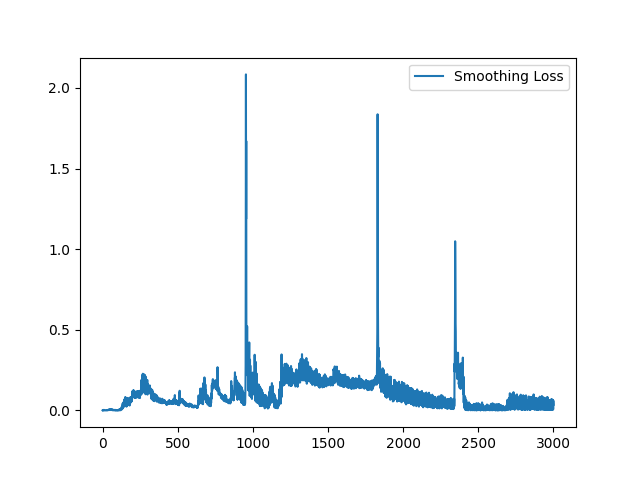
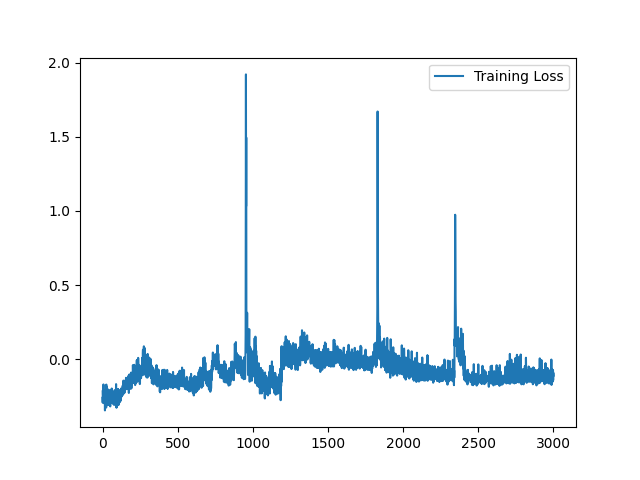
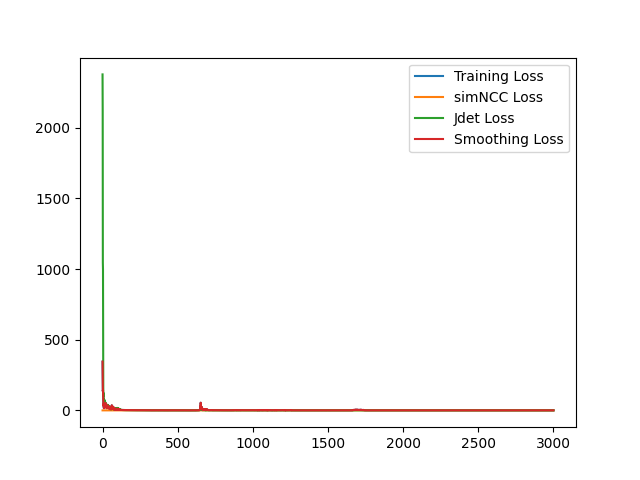
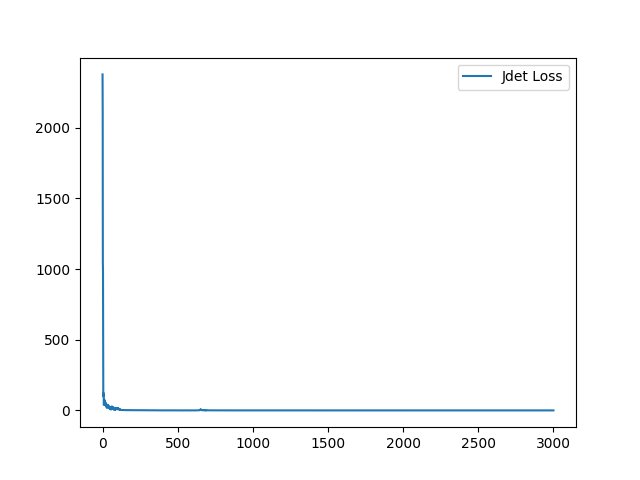
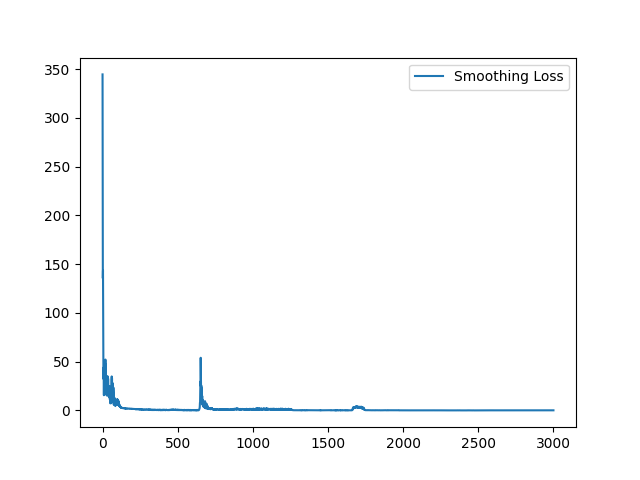
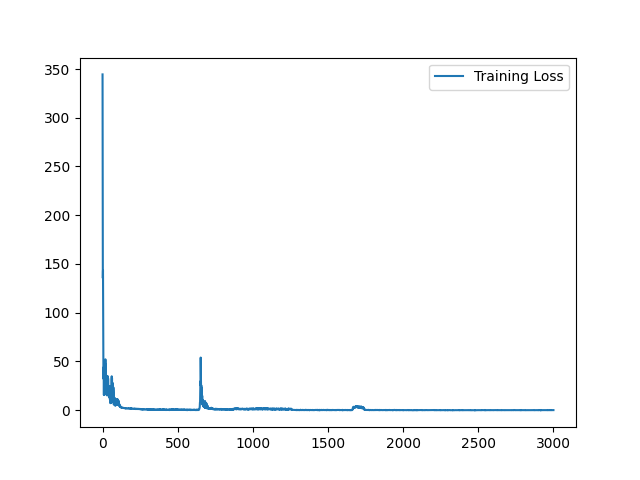
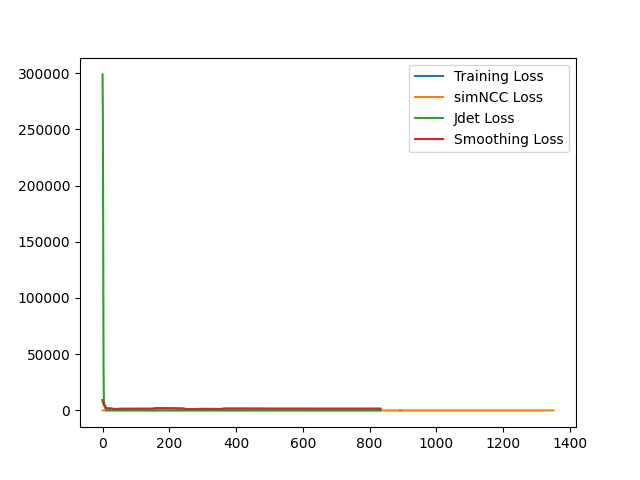
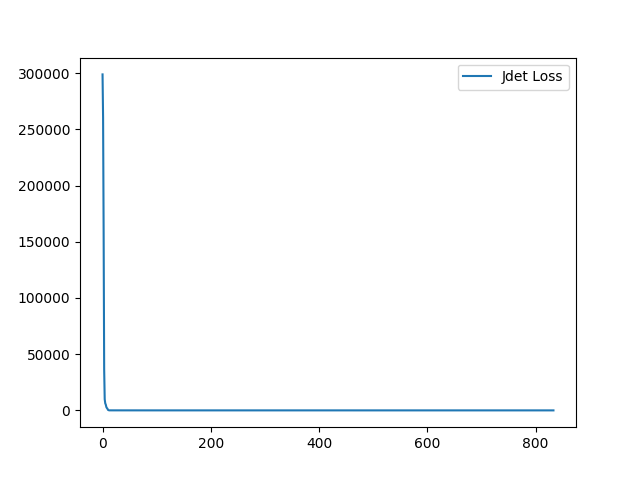
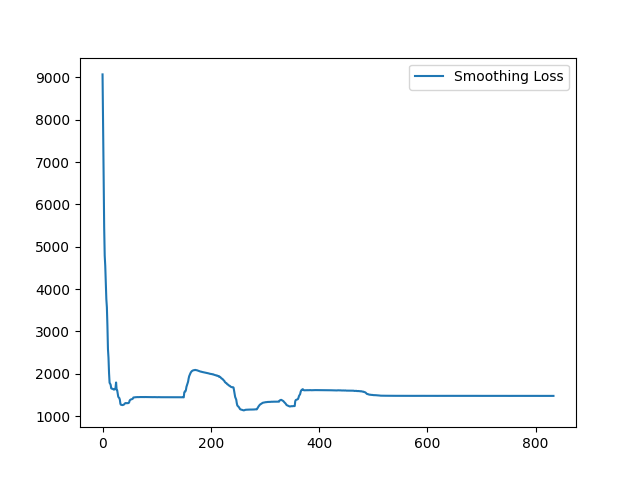
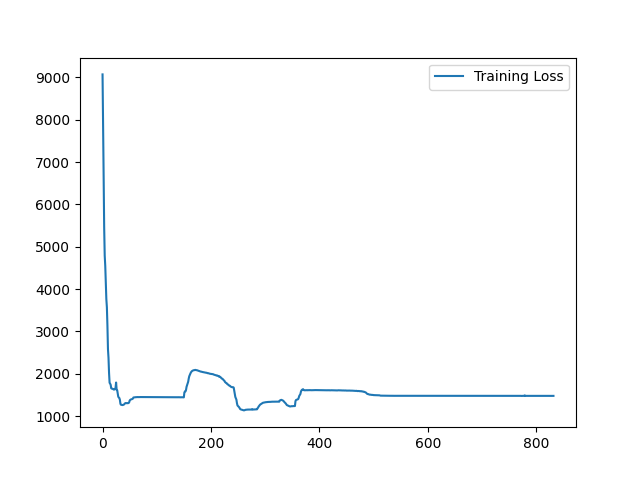
I have fixed the normalization method. Here is the sample training loss for your reference:
|
Just remember do not use zero-mean 1-std normalization with NCC function. Please use min-max normalization instead (i.e.: [0, 1].
Yes, it is normal at the lower level. At the higher level, the resulting deformation field will be more complex and the Jdet will not = 0 for method parameterized with displacement field. For the diffeomorphic version, Jdet will be closed to zero across all the level.
Yes, it is expected. Similar (less misalignment) input pair seems to yield a better loss value because it is easier to register. On the contrary, images pair with large appearance difference tends to yield higher loss value. To visualize the training loss, visualizing the loss value for each Epoch will help. |
|
@cwmok thanks for your patience and support.
This is what I am doing. As I understood, everything now is as expected. FOrgive me if I have many questions, since your paper is a good paper, my goal is to write a tutorial about it providing a colab notebook with some working cases.
As I understood, since the training loss is going up and down, we can not judge the performance until we test the model, right? More questions :)
|
No problem. And thanks a lot for your interest in our work. :)
Yes, like other deep learning applications, you need to implement your own validation code using validation data during training. In my paper, we use Dice score of the segmentation map to search for the best model during training (code is not provided as it is too customized).
In the paper, we didn't use any augmentation, which provides a fair comparison to other state-of-the-art methods. However, in the Learn2Reg challenge, we did use augmentation (i.e. affine augmentation and random horizontal flipping). Empirically, 3-5% gains in Dice score when augmentation is applied in a small dataset. |
|
These are the current training results, I noticed the following:
Since you already worked with the same dataset, could you please give some feedback to re-produce your result and explain why I am getting these results? level 1 training level 2 training level 3 training |
|
Hi @idhamari,
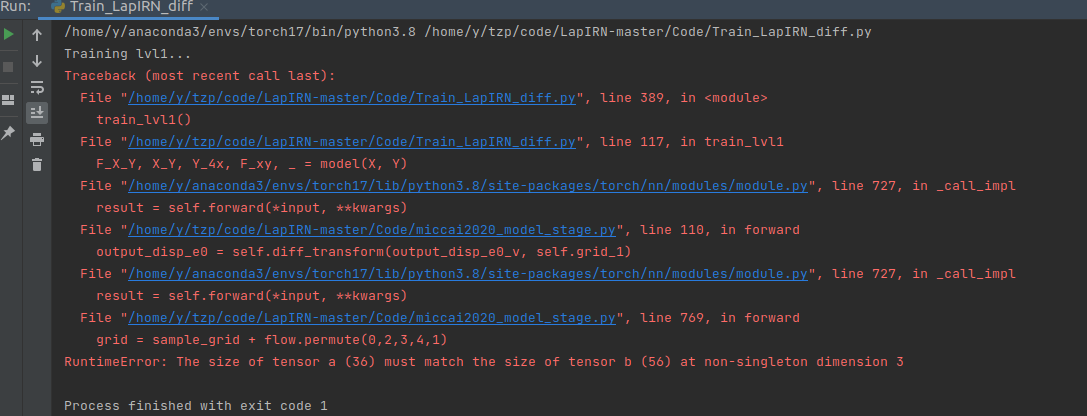
Debugging processObservation:
Conclusion:
I hope the debugging process and the conclusion help. Regards, |
|
Thanks a lot for your feedback, I really appreciate it. This step is not clear: apply the windowing with lower and upper bound set to -500 and 800" Shall I map normalize the image to [0,1] then map the values to [-500,800] range? this will conflict with "Making sure that the intensity value of the image's background is zero". I checked the paper in your link, it does not explain this step or provide a reference to explain it. |
No, windowing also known as grey-level mapping, contrast stretching, histogram modification or contrast enhancement. It is common in CT preprocessing pipeline as to locate the target anatomical structure for subsequent analysis. For more detail, you may refer to this website. To achieve this, one way is to apply the numpy clip function to the raw image and set the min and max to your desired HU, i.e.: [-500, 800] in this case, so that all the anatomical structures are clear for observation. At the last step, you have to normalize the preprocessed image to [0, 1] such that the background's intensity for each image is zero. |
|
@cwmok Should I change anything? |
|
@tzp123456 |
|
Hi @cwmok |
|
@tzp123456
The code was tested with GTX1080ti GPU and RTX titan GPU, which has 11GB and 24GB GPU memory, respectively. The default setting will consume around 10.8GB GPU memory. Therefore, if your GPU is used for display/OS/chrome, you will face the same error. In this case, try to lower the number of feature maps will help, e.g. set "--start_channel" to 6 instead of 7, it wouldn't make a big difference in terms of registration using OASIS dataset.
No, the moving images and fixed images are randomly selected during training, see Please open a new issue on my Github repository if you have further questions. Or if you have questions regarding @idhamari's tutorial, please contact him directly. |
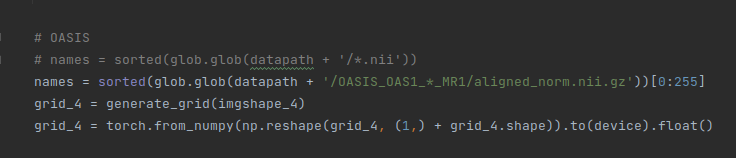
The updated code includes:
Future to do: