Multimodal image analysis pipeline combining MALDI mass spectrometry imaging, two-photon laser scanning microscopy (2PLSM), and H&E histology to characterize spatial heterogeneity in colorectal cancer tissue. Accompanies the publication in npj Imaging (2024).
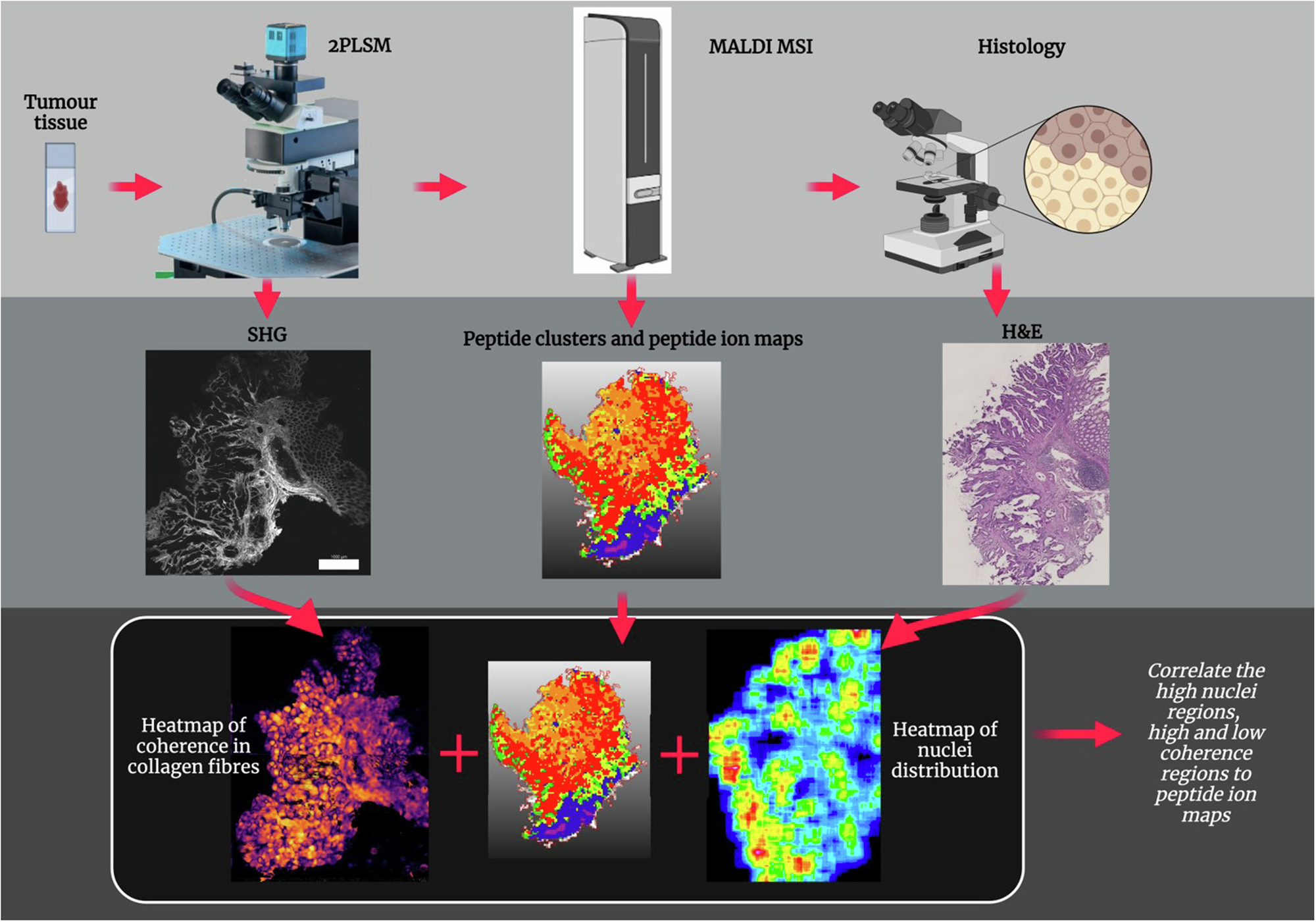
2PLSM (collagen texture)
- Structure tensor analysis — local fiber coherence (0 = chaotic, 1 = organised) and orientation (0–180°)
- Local density maps and fibrotic area quantification
- Orientation vector field visualisation
Histology (H&E)
- StarDist nuclei segmentation (deep learning,
2D_versatile_hemodel) - Kernel density estimation and nuclear morphometry (eccentricity, area)
- H&E stain normalisation (Ruifrok & Johnston colour deconvolution)
- Proximity network graphs and Voronoi tessellation of nuclei
MALDI-MSI
- Open-source pipeline replacing SCiLS Lab: imzML import, TIC normalisation, peak alignment
- ROC analysis per m/z, discriminative ion selection
- PCA and bisecting k-means spatial segmentation
- QuPath annotation import (GeoJSON)
git clone https://github.com/arora-bharti/MALDI-MSI_2PLSM_Histology.git
cd MALDI-MSI_2PLSM_Histology
python -m venv venv && source venv/bin/activate
pip install -r requirements.txtOr with Docker (no install required — reproduces the exact analysis environment):
docker build -t maldi-pipeline .
docker run -p 8501:8501 maldi-pipelinestreamlit run app/streamlit_app.py# Collagen texture analysis
python scripts/analyze_texture.py -i image.tif -o results/
# Nuclei segmentation
python scripts/segment_nuclei.py -i histology.tif -o results/| Notebook | Purpose |
|---|---|
Collagen_textureanalysis.ipynb |
Step-by-step 2PLSM texture analysis |
Nuclei_segmentation.ipynb |
Nuclei detection and density maps |
Low_high_coherence_percentage.ipynb |
Coherence statistics across images |
Orientation_vectorfield.ipynb |
Fibre orientation quiver overlays |
MALDI_analysis.ipynb |
Full MALDI-MSI pipeline |
pytest tests/ -vTests for TensorFlow/StarDist-dependent functions are skipped automatically if those libraries are not installed.
| Parameter | Description | Typical range |
|---|---|---|
filter_sigma |
Gaussian smoothing sigma | 1–5 |
local_sigma |
Structure tensor window | 5–20 |
local_density_kernel |
Kernel size for density | 10–30 |
| Parameter | Description | Default |
|---|---|---|
prob_thresh |
Detection probability threshold | 0.4 |
nms_thresh |
Non-maximum suppression threshold | 0.3 |
normalization_percentiles |
Input normalisation range | [25, 85] |
Texture analysis: *_filtered.tif, *_density.tif, *_coherence.tif, *_orientation.tif, Results_*.png, *.csv
Nuclei segmentation: labeled/*.tif, density/*.tif, circularity/*.tif, area/*.tif, *_mosaic.png, results/*.csv
@article{arora2024maldi,
title={MALDI imaging combined with two-photon microscopy reveals local differences in the heterogeneity of colorectal cancer},
author={Arora, Bharti and Kulkarni, Ajinkya and Markus, M. Andrea and Ramos-Gomes, Fernanda and Bohnenberger, Hanibal and Str{\"o}bel, Philipp and Alves, Frauke and Klein, Oliver},
journal={npj Imaging},
volume={2},
number={35},
year={2024},
doi={10.1038/s44303-024-00041-3}
}GNU Affero General Public License v3.0 — see LICENSE.
Bharti Arora and Ajinkya Kulkarni Max Planck Institute for Multidisciplinary Sciences, Göttingen, Germany
Funded by: EU Horizon 2020 Marie Sklodowska-Curie (No 857894 – CAST), Ministry for Science and Culture of Lower Saxony (ABA), Federal Ministry of Education and Research (MSTAR #031L0220A), BCRT and Berlin Institute of Health.
Repository structure
MALDI-MSI_2PLSM_Histology/
├── src/
│ ├── modules_2photon.py # 2PLSM texture analysis
│ ├── modules_histo.py # Histology / nuclei analysis
│ ├── modules_maldi.py # MALDI-MSI analysis
│ └── __init__.py
├── notebooks/ # Jupyter notebooks (one per modality)
├── scripts/
│ ├── analyze_texture.py # CLI for texture analysis
│ ├── segment_nuclei.py # CLI for nuclei segmentation
│ └── Merging.ijm # ImageJ macro for image overlay
├── app/
│ └── streamlit_app.py # Interactive web app
├── tests/ # pytest unit tests (52 tests, synthetic data)
├── sample_data/ # Place sample .tif images here
├── Dockerfile # Reproducible environment
├── requirements.txt
└── CITATION.cff


