,-,-----,
PIGPEN **** \ \ ),)`-'
<`--'> \ \`
/. . `-----,
OINC! > ('') , @~
`-._, ___ /
-|-|-|-|-|-|-|-| (( / (( / -|-|-|
|-|-|-|-|-|-|-|- ''' ''' -|-|-|-
-|-|-|-|-|-|-|-|-|-|-|-|-|-|-|-|-|
Pipeline for Identification
Of Guanosine Positions
Erroneously Notated
OINC-seq (Oxidation-Induced Nucleotide Conversion sequencing) is a sequencing technology that allows the direction of oxidative marks on RNA molecules. Because guanosine has the lowest redox potential of any of the ribonucleosides, it is the one most likely to be affected by oxidation. When this occurs, guanosine is turned into 8-oxoguanosine (8-OG) or further oxidized products. When reverse transcriptase encounters these products, it makes predictable errors in the resulting cDNA (see here). OINC-seq employs spatially restricted singlet oxygen radicals to oxidize RNAs at specific subcellular locations. The level of RNA oxidation detected for each RNA species is therefore a readout of the amount of that RNA species at that subcellular location.
To detect and quantify these conversions, we have created software called PIGPEN (Pipeline for Identification of Guanosine Positions Erroneously Notated).
PIGPEN starts with RNAseq fastq files. These files are aligned to the genome using STAR. Single and paired-end reads are supported, although paired-end reads are preferred (for reasons that will become clear later). To minimize the contribution of positions that appear as mutations due to non-ideal alignments, PIGPEN only considers uniquely aligned reads (mapping quality == 255). For now, it is required that paired-end reads be stranded, and that read 1 correspond to the sense strand. This is true for most, but not all, modern RNAseq library preparation protocols.
Uniquely aligned reads are then extracted and used to quantify transcript abundances using salmon. Posterior probabilities of transcript assignments are then derived using postmaster. STAR, salmon, and postmaster must be in the user's $PATH. All three of these preparatory steps can be easily and automatically done using alignAndQuant.py.
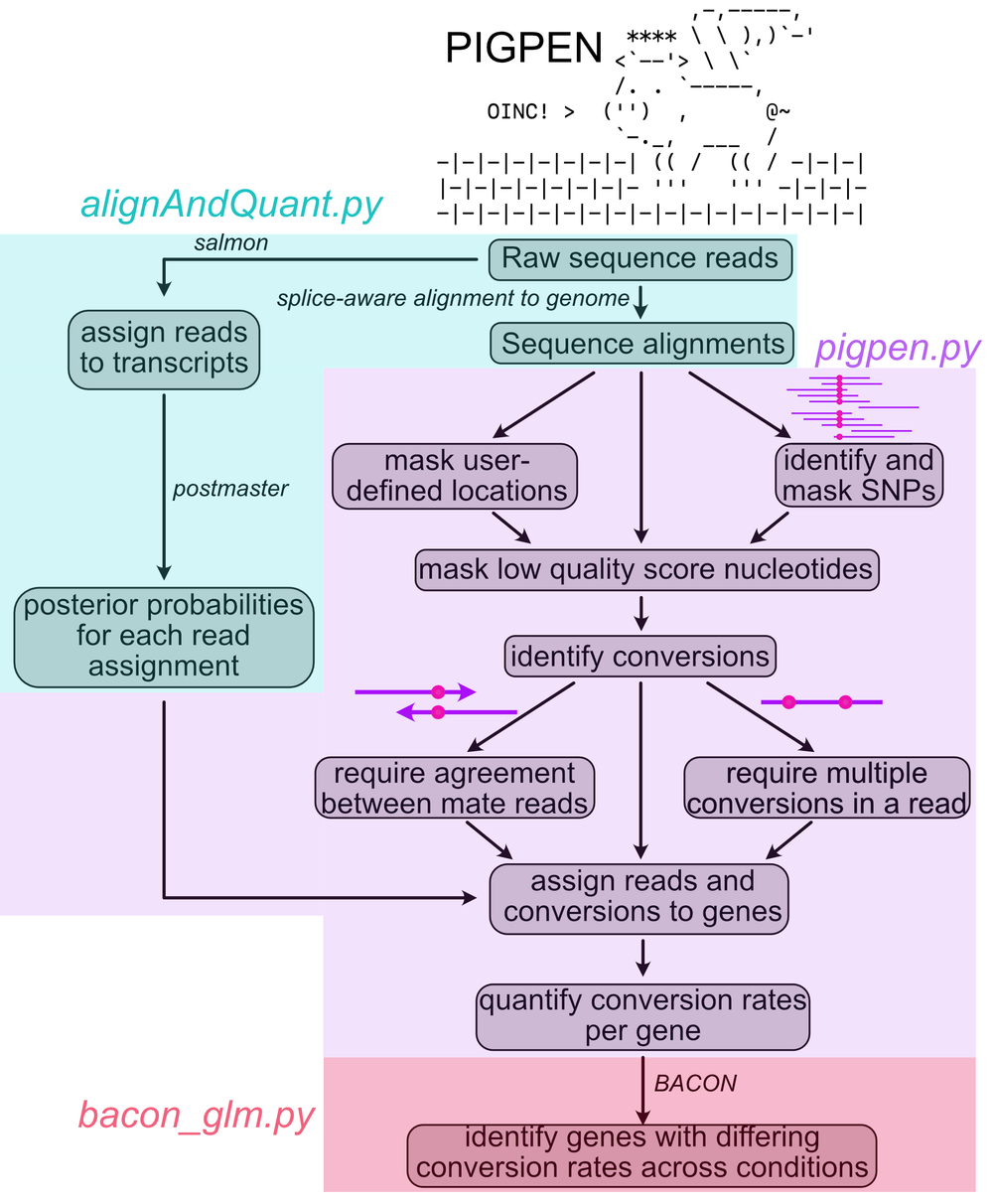
Following the creation of alignment files produced by STAR and postmaster as well as transcript quantifications produced by salmon, these files are then used by pigpen.py to identify nucleotide conversions, assign them to transcripts and genes, and then quantify the number of conversions in each gene. A graphical overview of the flow of PIGPEN is shown below.
PIGPEN has the following prerequisites:
- python >= 3.8
- samtools >= 1.15
- varscan >= 2.4.4
- bcftools >= 1.15
- pysam >= 0.19
- numpy >= 1.21
- pybedtools >= 0.9.0
- pandas >= 1.3.5
- bamtools >= 2.5.2
- salmon >= 1.9.0
- STAR >= 2.7.10
- gffutils >= 0.11.0
- umi_tools >= 1.1.0 (if UMI collapsing is desired)
- postmaster
Note: postmaster is a rust package. Installing it requires rust (which itself is installable using conda). Once rust is installed, use
cargo install --git https://github.com/COMBINE-lab/postmasterto install postmaster.
BACON has the following prerequisites:
- python >= 3.6
- statsmodels >= 0.13.2
- numpy >= 1.21
- rpy2 >= 3.4.5
- R >= 4.1
PIGPEN can be installed using bioconda using conda install -c bioconda pigpen. Following installation using conda, PIPGEN is accessible by calling pigpen, e.g. pigpen -h. Currently, postmaster must be installed separately afterward. This can be done using cargo install --git https://github.com/COMBINE-lab/postmaster. If PIPGEN was installed via conda, make sure to install postmaster in the same environment. #TODO: make bacon and alignAndQuant easily accessible after installation.
Alternatively, you can download PIGPEN directly from this repository. PIPGEN is python-based, but requires a number of extra modules as well as some R and Rust libraries. These are most easilty installed with conda. The necessary software is listed in pigpen_env.yaml. This configuration file can be provided to conda and has all the information needed to setup a PIGPEN-ready environment.
conda env create -f pigpen_env.yaml
This will create an environment called pigpen_env that contains all the necessary modules. To activate the environment, type
source activate pigpen_env
Uncompress the repository and move into the compressed directory. Install PIGPEN using
python setup.py install
Then to make sure you are ready to go, ask for the help options in the PIGPEN, BACON, and alignAndQuant scripts using
pigpen -h
bacon -h
alignAndQuant -h
If there are errors, one or more of the modules likely did not install properly. In that case, using an alternative package manager like pip may help. If you see no errors, you are good to go.
pigpen expects a particular directory structure for organization of STAR, salmon, and postmaster outputs. This is represented below.
workingdir
│
└───sample1
│ │
│ └───STAR
│ │ │ sample1Aligned.sortedByCoord.out.bam
│ │ │ sample1Aligned.sortedByCoord.out.bam.bai
│ │ │ ...
│ │
│ └───salmon
│ │ │ sample1.quant.sf
│ │ │ sample1.salmon.bam
│ │ │ ...
│ │
│ └───postmaster
│ │ │ sample1.postmaster.bam
│ │ │ sample1.postmaster.bam.bai
│ │ │ ...
│
└───sample2
│ │
│ └───STAR
│ │ │ sample2Aligned.sortedByCoord.out.bam
│ │ │ sample2Aligned.sortedByCoord.out.bam.bai
│ │ │ ...
│ │
│ └───salmon
│ │ │ sample2.quant.sf
│ │ │ sample2.salmon.bam
│ │ │ ...
│ │
│ └───postmaster
│ │ │ sample2.postmaster.bam
│ │ │ sample2.postmaster.bam.bai
│ │ │ ...
...
This structure can be automatically acheived by running alignAndQuant in workingdir once for each sample. Following this, the samples are ready to be analyzed with pigpen.
For example:
alignAndQuant --forwardreads reads.r1.fq.gz --reversereads reads.r2.fq.gz --nthreads 32 --STARindex <STARindex> --salmonindex <salmonindex> --samplename sample1
STARindex and salmonindex should be created according to the instructions for creating them found here and here.
Samples are then ready for analysis with pigpen.py. From workingdir, a comma-separated list of samples is supplied to --samplenames. In the example above, this would be --samplenames sample1,sample2. Optionally, a list of control samples are provided to --controlsamples. These should correspond to samples in which nucleotide conversions were not intentionally induced. They serve as controls for SNP identification (see below). They may be a subset of the samples provided to --samplenames.
8-OG-induced conversions are rare, and this rarity makes it imperative that contributions from conversions that are not due to oxidation are minimized. A major source of apparent conversions is SNPs. It is therefore advantageous to find and mask SNPs in the data.
PIGPEN performs this by using varscan to find SNP positions. These locations are then excluded from all future analyses. Varscan parameters are controled by the PIGPEN parameters --SNPcoverage and --SNPfreq that control the depth and frequency required to call a SNP. We recommend being aggressive with these parameters. We often set them to 20 and 0.2, respectively.
PIGPEN performs this SNP calling on control samples (--controlsamples) in which the intended oxidation did not occur. PIGPEN will use the union of all SNPs found in these files for masking. Whether or not to call SNPs at all (you probably should) is controlled by --useSNPs.
PIGPEN uses genome annotations to relate transcripts and genes. There are peculiarities to these annotations based on where they came from. Generally, PIGPEN prefers annotation files from either GENCODE or Ensembl. Tell PIGPEN where your annotation came from using the --gfftype flag.
PIGPEN then identifies conversions in reads. This can be done using multiple processors (--nproc). In order to minimize the effect of sequencing error, PIGPEN only considers positions for which the sequencing quality was at least 30. There are two important flags to consider here.
First, --onlyConsiderOverlap requires that the same conversion be observed in both reads of a mate pair. Positions interrogated by only one read are not considered. This can improve accuracy. True oxidation-induced conversions are rare. Rare enough that sequencing errors can cause a problem. Requiring that a conversion be present in both reads minimizes the effect of sequencing errors. If the fragment sizes for a library are especially large relative to the read length, the number of positions interrogated by both mates will be small.
Second, nConv sets the minimum number of G -> C / G -> T conversions in a read pair in order for those conversions to be recorded. The rationale here is again to reduce the contribution of background, non-oxidation-related conversions. Background conversions should be distributed relatively randomly across reads. However, due to the spatial nature of the oxidation reaction, oxidation-induced conversions should be more clustered into specific reads. Therefore, requiring at least two conversions can increase specificity. In practice, this works well if the data is very deep or concentrated on a small number of targets. When dealing with transcriptome-scale data, this flag often reduces the number of observed conversions to an unacceptably low level.
After PIGPEN calculates the number of converted and noncoverted nucleotides in each read pair, it intersects that data with the probabilistic transcript assignment for each read performed by salmon and postmaster. Conversions within read pair X are assigned proportionally to transcript Y according to the salmon/postmaster-calculated probability that read pair X originated from transcript Y. This transcript-level data is then collapsed to gene-level data according to the transcript/gene relationships found in --gff. Transcript IDs in --gff should match those in the fasta file used to make --salmonindex. The use of GENCODE annotations is recommended if possible. Alternatively, Ensembl annotations can be used. The source of the annotations should be supplied using the --gfftype flag.
We have observed that the overall rate of conversions (not just G -> T + G -> C, but all conversions) can vary signficantly from sample to sample, presumably due to a technical effect in library preparation. For this reason, PIGPEN calculates PORC (Proportion of Relevant Conversions) values. This is the log2 ratio of the relevant conversion rate ([G -> T + G -> C] / total number of reference G encountered) to the overall conversion rate (total number of all conversions / total number of positions interrogated). PORC therefore normalizes to the overall rate of conversions, removing this technical effect.
PIGPEN can use G -> T conversions, G -> C conversions, G deletions, or any combination when calculating PORC values. This behavior is controlled by supplying some or all of the options --use_g_t, --use_g_c, and --use_g_x, respectively.
The use of one read in a paired end sample for conversion quantification can be controlled using --use_read1 and --use_read2. To use both reads, supply both flags. --onlyConsiderOverlap requires the use of both reads. Importantly, both reads can still used for genomic alignment and transcript quantification.
To prevent specific genomic locations from being considered during conversion quantification, supply a bed file of these locations to --maskbed.
Output files are named <samplename>.pigpen.txt. These files contain the number of observed conversions for each gene as well as derived values like conversion rates and PORC values.
We could simply compare PORC values across conditions, but with that approach we lose information about the number of counts (conversions) that went into the PORC calculation.
For each gene, PIGPEN calculates the number of relevant conversions (G -> T + G -> C) as well as all other conversions encountered. Each gene therefore ends up with a 2x2 contingency table of the following form:
| converted | not converted | |
|---|---|---|
| G -> C or G -> T | a | b |
| other conversion | c | d |
We then want to compare groups (replicates) of contingency tables across conditions. BACON (Bioinformatic Analysis of the Conversion of Nucleotides) performs this comparison using a binomial linear mixed-effects model. Replicates are modeled as random effects.
full model = conversions ~ nucleotide + condition + nucleotide:condition + (1 | replicate)
null model = conversions ~ nucleotide + condition + (1 | replicate)
The two models are then compared using a likelihood ratio test.
As input, BACON takes a tab-delimited, headered file of the following form with one row per sample:
| file | sample | condition |
|---|---|---|
| /path/to/pigpen/output | sample name | condition ID |