Drug Response Omics association MAp (DROMA, 卓玛)
Visit the DROMA website at: https://droma01.github.io/DROMA/
Explore the interactive web interface with comprehensive documentation, component details, and live examples.
- 🌟 Overview
- 🎯 Key Features
- 🏗️ DROMA Ecosystem
- 📊 Dataset Overview
- 🚀 Quick Start
- 🔧 Installation
- 💻 Usage Examples
- 🌐 Web Interface
- 🤖 AI Integration
- 🖥️ Official Website
- 📖 Documentation
- 🤝 Contributing
- 📄 Citation
- 📧 Contact
- 📝 License
DROMA is a comprehensive precision oncology platform that integrates the world's largest collection of drug response and multi-omics datasets. It bridges the gap between cancer pharmacogenomics data and actionable insights through advanced analytics, AI-powered tools, and user-friendly interfaces.
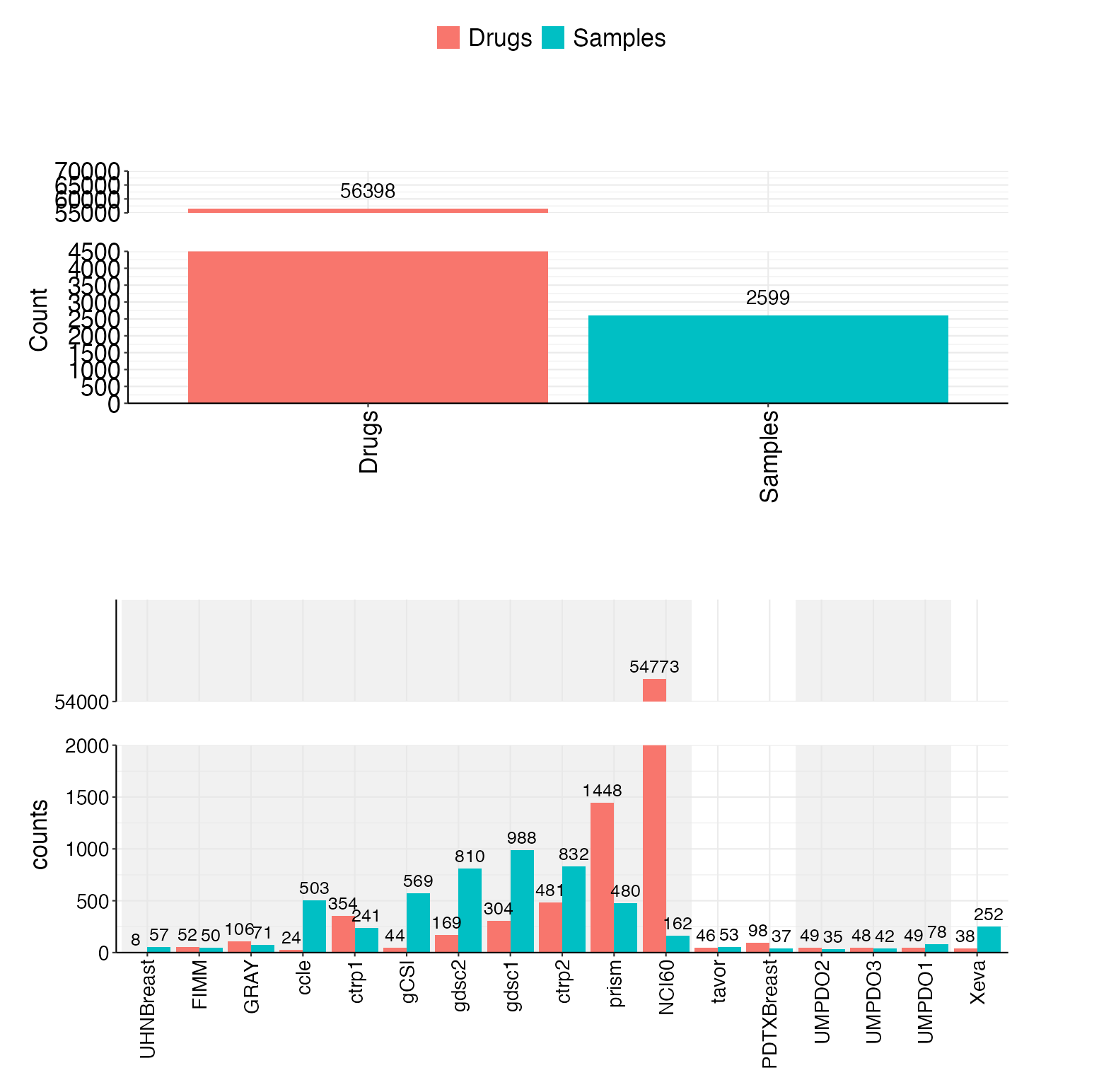
- 🗄️ Massive Dataset: 20 projects with 2,600+ samples and 56,000+ drugs - the world's largest open-source integrated drug sensitivity dataset
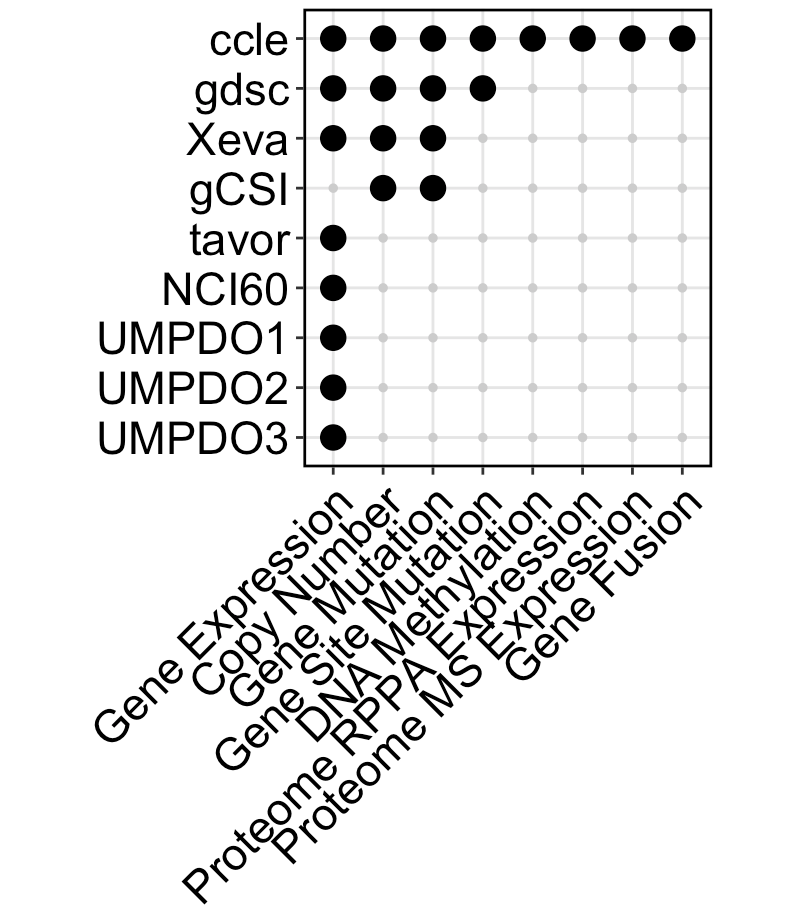
- 🧬 Multi-Omics Support: mRNA, CNV, mutations, methylation, proteomics, and more
- 🏥 Diverse Model Systems: Cell lines, PDOs (Patient-Derived Organoids), PDXs (Patient-Derived Xenografts), and PDCs (Patient-Derived Cells)
- 🤖 AI-Powered: Natural language interactions via MCP (Model Context Protocol) server
- 📊 Advanced Analytics: Meta-analysis, biomarker discovery, and statistical modeling
- 🌐 Web Interface: Interactive Shiny application for intuitive analysis
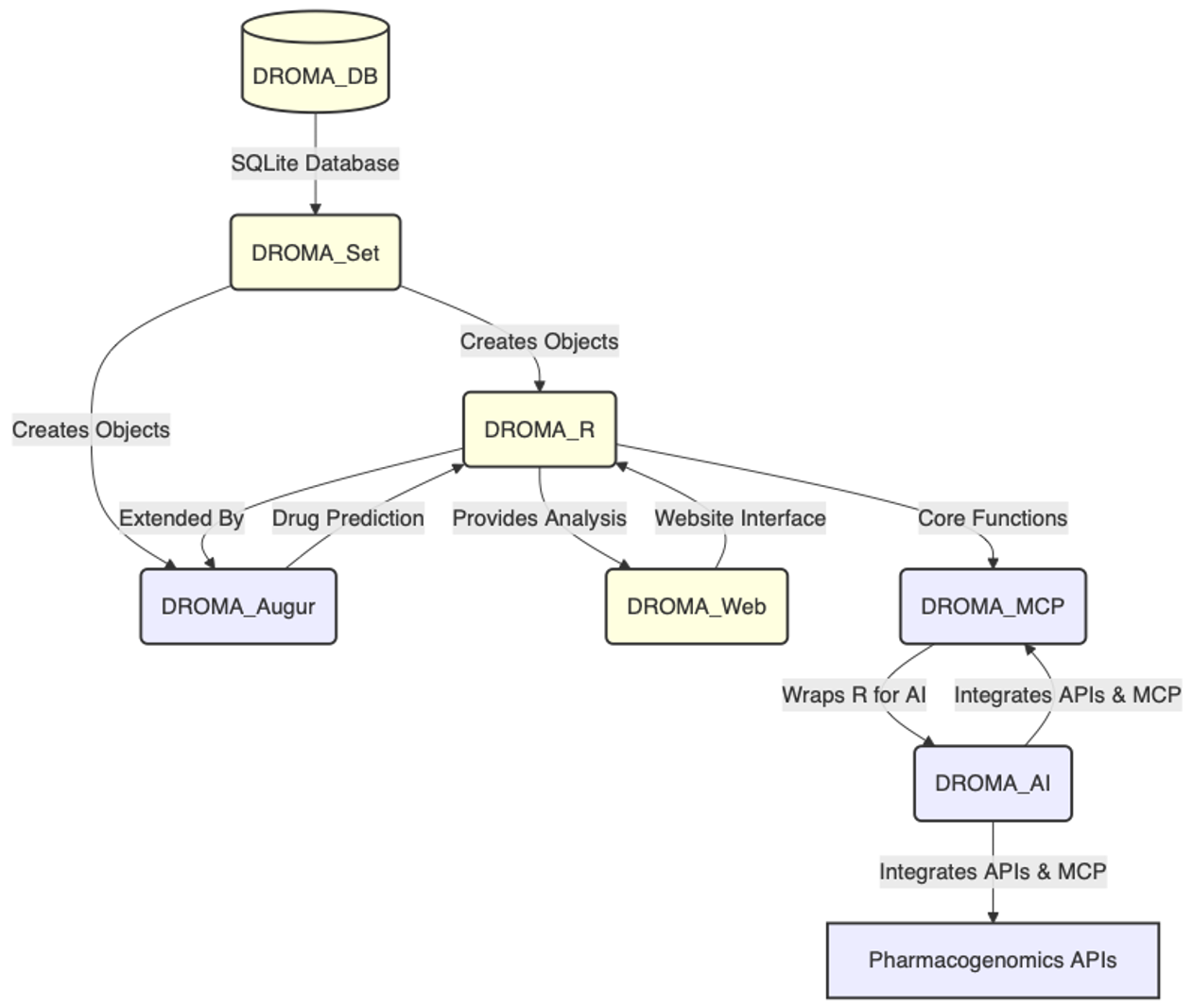
The DROMA platform consists of interconnected components that work together to provide a complete drug response analysis solution:
DROMA_DB - Database Foundation
- Purpose: SQLite database creation and management
- Features: Project-oriented structure, efficient querying, optimized indexing
- Content: 20 projects with comprehensive omics and drug response data
- Integration: Foundation for all other DROMA components
DROMA_Set - Data Management
- Purpose: R package for multi-project data management and analysis
- Features: S4 classes (DromaSet, MultiDromaSet), cross-project comparisons, sample overlap detection
- Key Classes:
DromaSet: Single-project analysisMultiDromaSet: Cross-project comparisons
- Capabilities: Flexible data loading, metadata management, database connectivity
DROMA_R - Advanced Analytics
- Purpose: Statistical analysis and visualization for drug-omics associations
- Features: Meta-analysis, batch processing, comprehensive visualization
- Methods: Spearman correlation, Wilcoxon tests, Cliff's Delta effect sizes
- Outputs: Forest plots, volcano plots, comparison visualizations
- Performance: Z-score normalization, parallel processing support
DROMA_Web - Web Application
- Purpose: Interactive Shiny web interface
- Features: Real-time analysis, dynamic visualization, user-friendly interface
- Modules: Drug feature analysis, batch analysis, drug-omics pairing
- Access: Browser-based, no installation required
DROMA_MCP - AI Interface
- Purpose: Model Context Protocol server for natural language interactions
- Features: AI assistant integration, natural language queries, automated analysis
- Capabilities: Dataset management, data loading, database exploration
- Integration: Works with ChatGPT, Claude, and other AI assistants
DROMA_Py - Python Access Layer
- Purpose: Python package for database operations and data access
- Features: Full DROMA ecosystem integration, Pythonic API, seamless R-Python bridge
- Capabilities: Database queries, data harmonization, batch processing, cross-platform compatibility
- Installation: Available on PyPI (
pip install droma-py) - Target Users: Python data scientists, bioinformaticians, computational biologists
- DROMA_AI: Multi-agent systems for automated analysis and hypothesis generation
- DROMA_Augur: Machine learning models for drug response prediction
Our comprehensive database includes:
| Metric | Count | Description |
|---|---|---|
| Projects | 20 | Independent research datasets |
| Samples | 2,600+ | Unique biological samples |
| Drugs | 56,000+ | Unique chemical compounds |
| Model Systems | 4 types | Cell lines, PDOs, PDXs, Clinical |
- Cell Lines (11 projects): CCLE, GDSC1, GDSC2, CTRP1, CTRP2, gCSI, NCI60, FIMM, GRAY, Prism, UHNBreast
- PDOs (5 projects): UMPDO1, UMPDO2, UMPDO3, HKPDO, LICOB
- PDCs (2 projects): PDTXBreast, Tavor
- PDXs (1 project): Xeva
- Clinical (1 project): CTR-DB (under development)
# Install DROMA.Set
devtools::install_github("mugpeng/DROMA_Set")
# Download data from Zenodo
# https://zenodo.org/records/15742800
# Connect database
library(DROMA.Set)
# Connect to your DROMA database
connectDROMADatabase("path/to/your/droma.sqlite")
# List available projects
projects <- listDROMAProjects()
print(projects)# Load dataset
gCSI <- createDromaSetFromDatabase("gCSI", "droma.sqlite")
# Load molecular profiles with normalization
gCSI <- loadMolecularProfilesNormalized(gCSI,
molecular_type = "mRNA",
features = "ABCB1")
# Load drug response
gCSI <- loadTreatmentResponseNormalized(gCSI, drugs = "Paclitaxel")
# Analyze drug-gene association
library(DROMA.R)
result <- analyzeDrugOmicPair(gCSI, "mRNA", "ABCB1", "Paclitaxel")# Install DROMA MCP
pip install droma-mcp
# Start AI server
droma-mcp run --db-path droma.sqlite
# Use with AI assistants
"Load CCLE dataset and analyze BRCA1 expression vs Tamoxifen response"# Install and use DROMA_Py
import droma_py as dp
# Connect to database
db = dp.DROMADatabase("droma.sqlite")
# Load and harmonize data
data = db.load_molecular_profiles("CCLE", "mRNA", features=["BRCA1", "TP53"])
harmonized = dp.harmonize_gene_names(data)
# Query drug response
drug_data = db.load_treatment_response("CCLE", drugs=["Tamoxifen"])Visit our interactive web application for browser-based analysis (contact for access).
- R ≥ 4.0.0
- Python ≥ 3.10 (for DROMA_MCP)
- SQLite database
# Install DROMA packages
devtools::install_github("mugpeng/DROMA_Set")
devtools::install_github("mugpeng/DROMA_R")# Install Python components
pip install droma-py # Python access layer
pip install droma-mcp # AI interface# Download from Zenodo (15.5 GB)
wget https://zenodo.org/records/15742800/files/droma.sqlite
unzip droma-data.zipEach component has comprehensive documentation:
- DROMA_DB Documentation: Database creation and management
- DROMA_Set Documentation: Data management and S4 classes
- DROMA_R Documentation: Statistical analysis and visualization
- DROMA_MCP Documentation: AI interface and natural language queries
Visit https://droma01.github.io/DROMA/ for the complete DROMA experience:
- Interactive Homepage: Animated DNA visualizations and ecosystem overview
- Component Documentation: Detailed information about each DROMA module
- Live Code Examples: Copy-to-clipboard functionality for quick setup
- Responsive Design: Optimized for desktop, tablet, and mobile devices
- Professional Theme: Scientific color palette with modern animations
- No Installation Required: Access DROMA information instantly in your browser
- Quick Navigation: Jump between components, documentation, and examples
- Mobile-Friendly: Full functionality on all device sizes
- Fast Loading: Optimized performance with modern web technologies
- Component Details: Deep-dive into DROMA_DB, DROMA_Set, DROMA_R, DROMA_Web, and DROMA_MCP
- Installation Guides: Step-by-step setup instructions for each component
- GitHub Integration: Direct links to all source repositories
- Data Access: Links to Zenodo datasets and documentation
- Identify genes associated with drug sensitivity across multiple datasets
- Perform meta-analysis to strengthen statistical power
- Validate findings across different model systems
- Explore drug response patterns across cancer types
- Identify compounds with similar response profiles
- Predict drug efficacy in new contexts
- Analyze patient-derived models (PDOs, PDXs)
- Correlate molecular profiles with treatment outcomes
- Develop personalized treatment strategies
- Compare drug responses across cell lines vs patient models
- Evaluate consistency of biomarkers across studies
- Integrate multi-omics data for comprehensive insights
All components work together through standardized interfaces and data formats.
- Optimized SQLite database with indexing
- Parallel processing for large-scale analysis
- Memory-efficient data handling
- Interactive plots and dashboards
- Publication-ready figures
- Customizable themes and layouts
- Filter by tumor type, data type, or specific features
- Cross-project sample matching
- Advanced statistical filtering
- Natural language queries
- Automated analysis workflows
- Intelligent data exploration
DROMA has been used for:
- Academic Research: Published in high-impact journals
- Drug Discovery: Industrial collaborations
- Clinical Translation: Biomarker validation studies
- Educational: Training in computational pharmacogenomics
- New: DROMA_MCP AI interface
- Enhanced: Multi-project analysis capabilities
- Added: Z-score normalization by default
- Improved: Performance optimizations
250620, I attended 11th Macau Symposium on Biomedical Sciences 2025 - 11th Macau Symposium on Biomedical Sciences 2025:
- Added: In vivo data (Xeva PDX dataset)
- Total: 20 datasets (11 cell line, 2 PDC, 5 PDO, 1 PDX, 1 clinical)
- Enhanced: Drug feature analysis modules
- Improved: Visualization and user experience
250319 UM PhD seminar:
- Launch: Interactive Shiny web application
- Added: PDO datasets (breast, colon, nasopharyngeal)
- Enhanced: Data harmonization and annotation
- Features: Tumor type and data type filtering
If you use DROMA in your research, please cite:
@article{li2024facilitating,
title={Facilitating integrative and personalized oncology omics analysis with UCSCXenaShiny},
author={Li, Shixiang and Peng, Yu and Chen, Miaozun and others},
journal={Communications Biology},
volume={7},
number={1},
pages={1200},
year={2024},
publisher={Nature Publishing Group},
doi={10.1038/s42003-024-06891-2}
}We welcome contributions to the DROMA ecosystem!
- Fork the relevant repository
- Create a feature branch
- Make your changes with tests
- Submit a pull request
- New datasets: Additional cancer pharmacogenomics studies
- Analysis methods: Novel statistical approaches
- Visualization: Enhanced plotting functions
- Documentation: Tutorials and examples
- Bug reports: Issues and feature requests
- Main Project: github.com/DROMA01/DROMA
- Database: github.com/mugpeng/DROMA_DB
- Data Management: github.com/mugpeng/DROMA_Set
- Analytics: github.com/mugpeng/DROMA_R
- Python Interface: github.com/mugpeng/DROMA_Py
- AI Interface: github.com/mugpeng/DROMA_MCP
- Data Repository: Zenodo Record
- Documentation: Component-specific README files
- Examples: Comprehensive usage examples in each repository
- Issues: Repository-specific GitHub Issues
- Discussions: GitHub Discussions
- Contact: yc47680@um.edu.mo
DROMA is licensed under the Mozilla Public License 2.0 (MPL-2.0). See LICENSE for details.
DROMA is developed by the precision oncology research team at the University of Macau. We thank all contributors, collaborators, and the broader cancer research community for their support and feedback.
Empowering precision medicine through comprehensive data integration and intelligent analysis