-
Notifications
You must be signed in to change notification settings - Fork 8
Expand file tree
/
Copy pathREADME.Rmd
More file actions
850 lines (693 loc) · 42.5 KB
/
README.Rmd
File metadata and controls
850 lines (693 loc) · 42.5 KB
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
326
327
328
329
330
331
332
333
334
335
336
337
338
339
340
341
342
343
344
345
346
347
348
349
350
351
352
353
354
355
356
357
358
359
360
361
362
363
364
365
366
367
368
369
370
371
372
373
374
375
376
377
378
379
380
381
382
383
384
385
386
387
388
389
390
391
392
393
394
395
396
397
398
399
400
401
402
403
404
405
406
407
408
409
410
411
412
413
414
415
416
417
418
419
420
421
422
423
424
425
426
427
428
429
430
431
432
433
434
435
436
437
438
439
440
441
442
443
444
445
446
447
448
449
450
451
452
453
454
455
456
457
458
459
460
461
462
463
464
465
466
467
468
469
470
471
472
473
474
475
476
477
478
479
480
481
482
483
484
485
486
487
488
489
490
491
492
493
494
495
496
497
498
499
500
501
502
503
504
505
506
507
508
509
510
511
512
513
514
515
516
517
518
519
520
521
522
523
524
525
526
527
528
529
530
531
532
533
534
535
536
537
538
539
540
541
542
543
544
545
546
547
548
549
550
551
552
553
554
555
556
557
558
559
560
561
562
563
564
565
566
567
568
569
570
571
572
573
574
575
576
577
578
579
580
581
582
583
584
585
586
587
588
589
590
591
592
593
594
595
596
597
598
599
600
601
602
603
604
605
606
607
608
609
610
611
612
613
614
615
616
617
618
619
620
621
622
623
624
625
626
627
628
629
630
631
632
633
634
635
636
637
638
639
640
641
642
643
644
645
646
647
648
649
650
651
652
653
654
655
656
657
658
659
660
661
662
663
664
665
666
667
668
669
670
671
672
673
674
675
676
677
678
679
680
681
682
683
684
685
686
687
688
689
690
691
692
693
694
695
696
697
698
699
700
701
702
703
704
705
706
707
708
709
710
711
712
713
714
715
716
717
718
719
720
721
722
723
724
725
726
727
728
729
730
731
732
733
734
735
736
737
738
739
740
741
742
743
744
745
746
747
748
749
750
751
752
753
754
755
756
757
758
759
760
761
762
763
764
765
766
767
768
769
770
771
772
773
774
775
776
777
778
779
780
781
782
783
784
785
786
787
788
789
790
791
792
793
794
795
796
797
798
799
800
801
802
803
804
805
806
807
808
809
810
811
812
813
814
815
816
817
818
819
820
821
822
823
824
825
826
827
828
829
830
831
832
833
834
835
836
837
838
839
840
841
842
843
844
845
846
847
848
849
850
---
output:
github_document:
toc: false
toc_depth: 1
editor_options:
chunk_output_type: console
---
<!-- README.md is generated from README.Rmd -->
```{r setup, include=FALSE}
knitr::opts_chunk$set(
collapse = TRUE,
eval = FALSE,
comment = "#>",
fig.path = "README-",
out.width = "100%"
)
```
<!-- badges: start -->
[](https://cran.r-project.org/package=ctrdata)
[](https://app.codecov.io/gh/rfhb/ctrdata)
[](https://github.com/rfhb/ctrdata/actions/workflows/check-standard-win-macos-linux.yaml)
<!-- badges: end -->
[Main features](#main-features) •
[References](#references) •
[Installation](#installation) •
[Overview](#overview-of-functions-in-ctrdata) •
[Data model](#data-model-of-ctrdata) •
[Databases](#databases-for-use-with-ctrdata) •
[Example workflow](#example-workflow) •
[Analysis across registers](#workflow-across-registers-example) •
[Tests, coverage](#tests-and-coverage) •
[Acknowledgements](#acknowledgements) •
[Future](#future-features)
# ctrdata for aggregating and analysing clinical trials
Package `ctrdata` provides functions for retrieving (downloading), aggregating and analysing clinical trials using information (structured protocol and result data, as well as documents) from public registers. It can be used with the
- EU Clinical Trials Register ("EUCTR", https://www.clinicaltrialsregister.eu/)
- EU Clinical Trials Information System ("CTIS", https://euclinicaltrials.eu/, [example](#workflow-ctis-example))
- ClinicalTrials.gov ("CTGOV2", https://clinicaltrials.gov/, [example](#workflow-ctgov-example))
- ISRCTN Registry ("ISRCTN", https://www.isrctn.com/)
Additional registers are being explored. The package facilitates investigating across registers the design and conduct of trials of interest, to describe their trends and availability for patients and to facilitate using their detailed results for research and meta-analyses. `ctrdata` is a package for the [R](https://www.r-project.org/) system, but other systems and tools can use the databases created with this package. This README was reviewed on 2026-04-18 for version 1.26.1.9000.
## Main features
* Trial information is easily found and downloaded: `ctrdata` generates queries for all registers or takes a user query in a register's web interface, then `ctrdata` retrieves in one go all protocol- and results-related trials data. A [script](#2-script-to-automatically-copy-users-query-from-web-browser) can automate using the query URL from all registers. Annotations can be made when downloading trials, [trial documents](#documents-example) and [historic versions](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html#historic-versions-of-trial-records-and-changes-in-sample-sizes) can be downloaded.
* Downloaded trial information is stored in a document-centric database, for fast and offline access. Information from different registers can be accumulated in a single collection. Uses `RSQLite`, `DuckDB`, `PostgreSQL` or `MongoDB`, see [Databases](#databases-that-can-be-used-with-ctrdata). Interactively browse through trial structure and data. Easily re-run a previous query to update trial records.
* For analyses, `ctrdata` functions suggest canonical [trial concepts](https://rfhb.github.io/ctrdata/reference/ctrdata-trial-concepts.html) to simplify analyses across registers, find synonyms of an active substance, identify unique (de-duplicated) trial records across all registers, merge and recode fields as well as easily access deeply-nested fields. Analyses can be done with `R` (see [vignette](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html)) and other systems, using the stored `JSON`-structured data.
Respect the registers' terms and conditions, see `ctrOpenSearchPagesInBrowser(copyright = TRUE)`. Please cite the package in any publication or work as follows:
<blockquote>
```{r, eval=TRUE, results='asis', echo=FALSE}
cat(rev(format(citation("ctrdata"), style = "text")), sep = " or <br/>")
```
</blockquote>
## References
An introduction to the package, together with worked examples and technical explanations is in:
* Herold R. Aggregating and analysing clinical trials data from multiple public registers using R package ctrdata. Research Synthesis Methods. 2026;17(3):624-656. [doi:10.1017/rsm.2025.10061](https://doi.org/10.1017/rsm.2025.10061)
Package `ctrdata` has been used for unpublished works and these publications:
- Jong et al. (2025) Experiences with Low-Intervention Clinical Trials -- the New Category under the European Union Clinical Trials Regulation. Clinical Trials <https://doi.org/10.1177/17407745241309293>
- Lopez-Rey et al. (2025) Use of Bayesian Approaches in Oncology Clinical Trials: A Cross-Sectional Analysis’. Frontiers in Pharmacology <https://doi.org/10.3389/fphar.2025.1548997>
- Russek et al. (2025) Supplementing Single-Arm Trials with External Control Arms—Evaluation of German Real-World Data. Clinical Pharmacology & Therapeutics <https://doi.org/10.1002/cpt.3684>
- Clinical Studies Sweden (2025) National Summary of Clinical Trials in Human Medicines Based on CTIS Data [link](https://www.kliniskastudier.se/english/news-archive/news-archive/2025-11-27-national-summary-of-clinical-trials-in-human-medicines-based-on-ctis-data)
- Alzheimer’s disease Horizon Scanning Report (2024) [link](https://www.ema.europa.eu/en/documents/report/alzheimers-disease-eu-horizon-scanning-report_en.pdf)
- Kundu et al. (2024) Analysis of Factors Influencing Enrollment Success in Hematology Malignancy Cancer Clinical Trials (2008-2023). Blood Meeting Abstracts <https://doi.org/10.1182/blood-2024-207446>
- Sood et al. (2022) Managing the evidence infodemic: Automation approaches used for developing NICE COVID-19 living guidelines. <https://doi.org/10.1101/2022.06.13.22276242>
- Lasch et al. (2022) The Impact of COVID‐19 on the Initiation of Clinical Trials in Europe and the United States. <https://doi.org/10.1002/cpt.2534>
- Blog post (2018) Innovation coming to paediatric research <https://paediatricdata.eu/innovation-coming-to-paediatric-research/>
- Cancer Research UK (2017) The impact of collaboration: The value of UK medical research to EU science and health. [link](https://www.ukri.org/wp-content/uploads/2023/02/MRC-150223-Theimpactofcollaboraton-ThevalueofUK-medicalresearchtoEUscienceandhealth.pdf)
- EMA (2017) Results of juvenile animal studies (JAS) and impact on anti-cancer medicine development and use in children [PDF file, p 34](https://www.ema.europa.eu/en/documents/scientific-guideline/results-juvenile-animal-studies-jas-and-impact-anti-cancer-medicine-development-and-use-children_en.pdf#page=34)
<!--
```{r}
# find publications using ctrdata
utils::browseURL("https://scholar.google.com/scholar?hl=en&as_sdt=0,5&q=%22ctrdata%22")
# Ferreira, Francisca Santos (2024)
# Reporte de redes de investigação nos ensaios clínicos pediátricos registados no EudraCT
# https://repositorio.ulisboa.pt/handle/10400.5/101030
# ctrdata is mentioned but not used in
# Sciannameo et al. (2024)
# Information Extraction from Medical Case Reports Using OpenAI InstructGPT.
# Computer Methods and Programs in Biomedicine
# [https://doi.org/10.1016/j.cmpb.2024.108326](https://doi.org/10.1016/j.cmpb.2024.108326)
```
-->
## Installation
### 1. Install package `ctrdata` in R
Package `ctrdata` is [on CRAN](https://cran.r-project.org/package=ctrdata) and [on GitHub](https://github.com/rfhb/ctrdata). Within [R](https://www.r-project.org/), use the following commands to install package `ctrdata`:
```{r install_ctrdata, eval=FALSE}
# Install CRAN version:
install.packages("ctrdata")
# Alternatively, install development version:
install.packages("devtools")
remotes::install_github("rfhb/ctrdata", build_vignettes = TRUE)
```
These commands also install the package's dependencies (`jsonlite`, `httr2`, `xml2`, `nodbi`, `stringi`, `lubridate`, `jqr`, `dplyr`, `zip`, `readr`, `rlang`, `htmlwidgets`, `stringdist` and `V8`).
### 2. Script to automatically copy user's query from web browser
Optional; works with all registers supported by `ctrdata` and is recommended for CTIS so that its URL in the web browser reflects the user's parameters for querying this register.
In the web browser, install the [Tampermonkey browser extension](https://www.tampermonkey.net/), click on "New user script" and then on "Tools", enter into "Import from URL" this URL: [`https://raw.githubusercontent.com/rfhb/ctrdata/master/tools/ctrdataURLcopier.js`](https://raw.githubusercontent.com/rfhb/ctrdata/master/tools/ctrdataURLcopier.js) and then click on "Install".
The browser extension can be disabled and enabled by the user. When enabled, the URLs to all user's queries in the registers are automatically copied to the clipboard and can be pasted into the `queryterm = ...` parameter of function [ctrLoadQueryIntoDb()](https://rfhb.github.io/ctrdata/reference/ctrLoadQueryIntoDb.html).
Additionally, this script retrieves results for `CTIS` when opening search URLs such as [`https://euclinicaltrials.eu/ctis-public/search#searchCriteria={"status":[3,4]}`](<https://euclinicaltrials.eu/ctis-public/search#searchCriteria={"status":[3,4]}>). After changing the URL in the browser, a "Reload page" is needed to conduct the search and show results.
## Overview of functions in `ctrdata`
Selected functions are listed in the approximate order of use in a user's workflow (in bold, main functions). See all functions in the [package documentation overview](https://rfhb.github.io/ctrdata/reference/index.html).
Function name | Function purpose
---------------------------- | --------------------------------------------
`ctrGenerateQueries()` | From simple user parameters, generates queries for each register to find trials of interest
`ctrOpenSearchPagesInBrowser()` | Open search pages of registers or execute search in web browser
`ctrLoadQueryIntoDb()` | Retrieve (download) or update, and annotate, information on trials from a register and store in a collection in a database
`ctrShowOneTrial()` | Show full structure and all data of a trial, interactively select fields of interest for `dbGetFieldsIntoDf()`
`dbFindFields()` | Find names of variables (fields) in the collection
`dbGetFieldsIntoDf()` | Create a data frame (or tibble) from trial records in the database with specific fields and trial concepts of interest calculated across registered, see [trial concepts](https://rfhb.github.io/ctrdata/reference/ctrdata-trial-concepts.html) for 20 trial concepts available
`dfTrials2Long()` | Transform the data.frame from `dbGetFieldsIntoDf()` into a long name-value data.frame, including deeply nested fields
`dfName2Value()` | From a long name-value data.frame, extract values for variables (fields) of interest (e.g., endpoints)
## Data model of `ctrdata`
Package `ctrdata` uses the data models that are implicit in data as retrieved from the different registers. No mapping is provided for any register's data model to a putative target data model. The reasons include that registers' data models are continually evolving over time, that only few data fields have similar values and meaning between registers, and that the retrieved public data may not correspond to the registers' internal data model. The structure of data for a specific trial can interactively be inspected and searched using function, see the section [below](#workflow-data-model).
Thus, the handling of data from different models of registers is to be done at the time of analysis. This approach allows a high level of flexibility, transparency and reproducibility. To support analyses, `ctrdata` (from version 1.21.0) provides functions that calculate concepts of clinical trials across registers, which are commonly used in analyses, such as start dates, age groups and statistical tests of results. See [help(ctrdata-trial-concepts)](https://rfhb.github.io/ctrdata/reference/ctrdata-trial-concepts.html) and the section [Analysis across trials](#workflow-across-registers-example) in the example workflow below. For further analyses, see examples of function [dfMergeVariablesRelevel()](https://rfhb.github.io/ctrdata/reference/dfMergeVariablesRelevel.html) on how to align related fields from different registers for a joint analysis.
In any of the [databases](https://rfhb.github.io/ctrdata/index.html#databases-for-use-with-ctrdata), one clinical trial is one document, corresponding to one row in a `SQLite`, `PostgreSQL` or `DuckDB` table, and to one document in a `MongoDB` collection. These `NoSQL` backends allow documents to have different structures, which is used here to accommodate the different models of data retrieved from the registers. Package `ctrdata` stores in every such document:
- field `_id` with the trial identification as provided by the register from which it was retrieved
- field `ctrname` with the name of the register (`EUCTR`, `CTGOV`, `CTGOV2`, `ISRCTN`, `CTIS`) from which that trial was retrieved
- field `record_last_import` with the date and time when that document was last updated using `ctrLoadQueryIntoDb()`
- only for `CTGOV2` and `CTIS`: object `history` with a historic version of the trial and with `history_version`, which contains the fields `version_number` (starting from 1) and `version_date`
- all original fields as provided by the register for that trial (see example in [vignette](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html#historic-versions-of-trial-records-and-changes-in-sample-sizes))
For visualising the data structure for a trial, see this
[vignette section](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html#analysing-nested-fields-such-as-trial-results).
## Databases for use with `ctrdata`
Package `ctrdata` retrieves trial data and stores it in a database collection, which has to be given as a connection object to parameter `con` for several `ctrdata` functions. This connection object is created almost identically for the four database backends supported by `ctrdata`, as shown in the table. For a speed comparison, see the [nodbi documentation](https://github.com/ropensci/nodbi#benchmark).
Besides `ctrdata` functions below, such a connection object can be used with functions of other packages, such as `nodbi` (see last row in table) or, in case of MongoDB as database backend, `mongolite` (see vignettes).
Purpose | Function call
-------------------- | --------------------
Create **SQLite** database connection | `dbc <- nodbi::src_sqlite(dbname = "name_of_my_database", collection = "name_of_my_collection")`
Create **DuckDB** database connection | `dbc <- nodbi::src_duckdb(dbdir = "name_of_my_database", collection = "name_of_my_collection")`
Create **MongoDB** database connection | `dbc <- nodbi::src_mongo(db = "name_of_my_database", collection = "name_of_my_collection")`
Create **PostgreSQL** database connection | `dbc <- nodbi::src_postgres(dbname = "name_of_my_database"); dbc[["collection"]] <- "name_of_my_collection"`
Use connection with `ctrdata` functions | `ctrdata::{ctrLoadQueryIntoDb, dbQueryHistory, dbFindIdsUniqueTrials, dbFindFields, dbGetFieldsIntoDf}(con = dbc, ...)`
Use connection with `nodbi` functions | e.g., `nodbi::docdb_query(src = dbc, key = dbc$collection, ...)`
## Vignettes
- [Install R package ctrdata](https://rfhb.github.io/ctrdata/articles/ctrdata_install.html)
- [Retrieve clinical trial information](https://rfhb.github.io/ctrdata/articles/ctrdata_retrieve.html)
- [Summarise and analyse clinical trial information](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html)
## Example workflow
The aim of the example is to download protocol-related trial information and tabulate the trials' status of conduct.
* Attach package `ctrdata`:
```{r load_ctrdata}
library(ctrdata)
```
* See help to get started with `ctrdata`:
```{r help_package}
help("ctrdata")
```
* Information on trial registers, their contents and how they can be used with `ctrdata` (last updated 2026-01-11):
```{r help_registers}
help("ctrdata-registers")
```
* Open registers' advanced search pages in browser:
```{r open_searchpages}
ctrOpenSearchPagesInBrowser()
# Please review and respect register copyrights:
ctrOpenSearchPagesInBrowser(copyright = TRUE)
```
* Adjust search parameters and execute search in browser
* When trials of interest are listed in browser, _copy the address from the browser's address bar to the clipboard_
* Search used in this example: https://www.clinicaltrialsregister.eu/ctr-search/search?query=neuroblastoma&phase=phase-two&age=children#tabs
* Get address from clipboard:
```{r get_clipboard}
q <- ctrGetQueryUrl()
# * Using clipboard content as register query URL: https://www.clinicaltrialsregister.eu/ctr-search/search?query=neuroblastoma&phase=phase-two&age=children
# * Found search query from EUCTR: query=neuroblastoma&phase=phase-two&age=children
q
# query-term query-register
# 1 query=neuroblastoma&phase=phase-two&age=children EUCTR
```
Queries in the trial registers can automatically copied to the clipboard (including for "CTIS", where the URL otherwise does not show the user's query), see [here](#2-script-to-automatically-copy-users-query-from-web-browser).
* Retrieve protocol-related information, transform and save to database:
```{r get_url, include=FALSE}
q <- "https://www.clinicaltrialsregister.eu/ctr-search/search?query=neuroblastoma&phase=phase-two&age=children"
ctrOpenSearchPagesInBrowser(q)
```
For loading the trial information, first a database collection is specified, using `nodbi` (see above for how to specify `PostgreSQL`, `RSQlite`, `DuckDB` or `MongoDB` as backend, see section [Databases](#databases-that-can-be-used-with-ctrdata)):
```{r create_db}
# Connect to (or create) an SQLite database
# stored in a file on the local system:
db <- nodbi::src_sqlite(
dbname = "database_name.sql",
collection = "collection_name"
)
```
Then, the trial information is retrieved and loaded into the collection:
```{r load_euctr}
# Retrieve trials from public register EUCTR,
# both protocol- and results-related data:
ctrLoadQueryIntoDb(
queryterm = q,
euctrresults = TRUE,
euctrprotocolsall = FALSE, # new since 2025-07-20, loads single
# instead of all available country versions of a trial in EUCTR
con = db
)
# * Found search query from EUCTR: query=neuroblastoma&phase=phase-two&age=children
# * Checking trials in EUCTR, found 73 trials
# - Downloading in 4 batch(es) (20 trials each; estimate: 5 s)...
# - Downloading 73 records of 73 trials (estimate: 4 s)...
# - Converting to NDJSON (estimate: 0.2 s)...
# - Importing records into database...
# = Imported or updated 73 records on 73 trial(s)
# * Checking results if available from EUCTR for 73 trials:
# - Downloading results...
# - Extracting results (. = data, F = file[s] and data, x = none): . . . F . . .
# F . . F . . . . . . . F . F F F . . . . . . F . . . .
# - Data found for 34 trials
# - Converting to NDJSON (estimate: 1 s)...
# - Importing 34 results into database (may take some time)...
# - Results history: not retrieved (euctrresultshistory = FALSE)
# = Imported or updated results for 34 trials
# No history found in expected format.
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 73
```
Under the hood, plain text from EUCTR and XML files from EUCTR, CTGOV, ISRCTN are converted using Javascript via `V8` in `R` into `NDJSON`, which is imported into the database collection.
* Easily generate queries for each register and add records from several registers into the same collection
The same parameters can be used to ask `ctrdata` to generate search queries that apply to each register, for opening the web interfaces and for loading the trial data into the collection:
```{r load_trials}
# Generate queries for each register
queries <- ctrGenerateQueries(
condition = "neuroblastoma",
recruitment = "completed",
phase = "phase 2",
population = "P"
)
queries
# EUCTR
# "https://www.clinicaltrialsregister.eu/ctr-search/search?query=neuroblastoma&phase=phase-two&age=children&age=adolescent&age=infant-and-toddler&age=newborn&age=preterm-new-born-infants&age=under-18&status=completed"
# ISRCTN
# "https://www.isrctn.com/search?&q=&filters=condition:neuroblastoma,phase:Phase II,ageRange:Child,trialStatus:completed,primaryStudyDesign:Interventional"
# CTGOV2
# "https://clinicaltrials.gov/search?cond=neuroblastoma&intr=Drug OR Biological&term=AREA[DesignPrimaryPurpose](DIAGNOSTIC OR PREVENTION OR TREATMENT)&aggFilters=phase:2,ages:child,status:com,studyType:int"
# CTGOV2expert
# "https://clinicaltrials.gov/expert-search?term=AREA[ConditionSearch]\"neuroblastoma\" AND (AREA[Phase]\"PHASE2\") AND (AREA[StdAge]\"CHILD\") AND (AREA[OverallStatus]\"COMPLETED\") AND (AREA[StudyType]INTERVENTIONAL) AND (AREA[DesignPrimaryPurpose](DIAGNOSTIC OR PREVENTION OR TREATMENT)) AND (AREA[InterventionSearch](DRUG OR BIOLOGICAL))"
# CTIS
# "https://euclinicaltrials.eu/ctis-public/search#searchCriteria={\"medicalCondition\":\"neuroblastoma\",\"trialPhaseCode\":[4],\"ageGroupCode\":[2],\"status\":[5,8]}"
# Open queries in registers' web interfaces
# Note the regular and expert CTGOV2 query
sapply(queries, ctrOpenSearchPagesInBrowser)
# Load all queries into database collection
result <- lapply(queries, ctrLoadQueryIntoDb, con = db)
sapply(result, "[[", "n")
# EUCTR ISRCTN CTGOV2 CTGOV2expert CTIS
# 180 0 105 105 2
```
* Analyse
Tabulate the status of trials that are part of an agreed paediatric development program (paediatric investigation plan, PIP). `ctrdata` functions return a data.frame (or a tibble, if package `tibble` is loaded).
```{r analyse_pips}
# Get all records that have values in the fields of interest:
result <- dbGetFieldsIntoDf(
# Field of interest
fields = c("a7_trial_is_part_of_a_paediatric_investigation_plan"),
# Trial concepts calculated across registers
calculate = c("f.statusRecruitment", "f.isUniqueTrial"),
con = db
)
# To review trial concepts details, call 'help("ctrdata-trial-concepts")'
# Querying database (16 fields)...
# Searching for duplicate trials...
# - Getting all trial identifiers (may take some time), 316 found in collection
# - Finding duplicates among registers' and sponsor ids...
# - 114 EUCTR _id were not preferred EU Member State record for 67 trials
# - Unique are 0 / 105 / 0 / 60 / 0 records from CTGOV / CTGOV2 / CTIS / EUCTR / ISRCTN
# = Returning keys (_id) of 165 records in collection "collection_name"
# Tabulate the clinical trial information of interest
with(
result[result$.isUniqueTrial, ],
table(
.statusRecruitment,
a7_trial_is_part_of_a_paediatric_investigation_plan
)
)
# a7_trial_is_part_of_a_paediatric_investigation_plan
# .statusRecruitment FALSE TRUE
# ongoing 4 3
# completed 11 6
# ended early 7 3
# other 8 2
```
<div id="workflow-ctgov-example"></div>
* Queries to CTGOV and CTGOV2
The new website and API introduced in July 2023 (https://clinicaltrials.gov/) is supported by `ctrdata` since mid-2023 and identified in `ctrdata` as `CTGOV2`.
On 2024-06-25, `CTGOV` retired the classic website and API used by `ctrdata` since 2015. To support users, `ctrdata` automatically translates and redirects queries to the current website. This helps with automatically updating previously loaded queries (`ctrLoadQueryIntoDb(querytoupdate = <n>)`), manually migrating queries and reproducible work on clinical trials information. Going forward, users are recommended to change to use `CTGOV2` queries.
As regards study data, important differences exist between field names and contents of information retrieved using `CTGOV` or `CTGOV2`; see the [schema for study protocols in `CTGOV`](https://cdn.clinicaltrials.gov/documents/xsd/prs/ProtocolRecordSchema.xsd), the [schema for study results](https://cdn.clinicaltrials.gov/documents/xsd/prs/RRSUploadSchema.xsd) and the [Study Data Structure for `CTGOV2`](https://clinicaltrials.gov/data-api/about-api/study-data-structure). For more details, call `help("ctrdata-registers")`. This is one of the reasons why `ctrdata` handles the situation as if these were two different registers and will continue to identify the current API as `register = "CTGOV2"`, to support the analysis stage.
Note that loading trials with `ctrdata` overwrites the previous record with `CTGOV2` data, whether the previous record was retrieved using `CTGOV` or `CTGOV2` queries.
Example using a CTGOV query:
```{r load_ctgov}
# CTGOV search query URL
q <- "https://classic.clinicaltrials.gov/ct2/results?cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug"
# Open old URL (CTGOV) in current website (CTGOV2):
ctrOpenSearchPagesInBrowser(q)
# Since 2024-06-25, the classic CTGOV servers are no longer available. Package ctrdata has translated the classic CTGOV query URL from this call of function ctrLoadQueryIntoDb(queryterm = ...) into a query URL that works with the current CTGOV2. This is printed below and is also part of the return value of this function, ctrLoadQueryIntoDb(...)$url. This URL can be used with ctrdata functions. Note that the fields and data schema of trials differ between CTGOV and CTGOV2.
#
# Replace this URL:
#
# https://classic.clinicaltrials.gov/ct2/results?cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug
#
# with this URL:
#
# https://clinicaltrials.gov/search?cond=neuroblastoma&intr=Drug&aggFilters=ages:child,results:with,status:com
#
# * Found search query from CTGOV2: cond=neuroblastoma&intr=Drug&aggFilters=ages:child,results:with,status:com
# [1] "https://clinicaltrials.gov/search?cond=neuroblastoma&intr=Drug&aggFilters=ages:child,results:with,status:com"
# Count trials
ctrLoadQueryIntoDb(
queryterm = q,
con = db,
only.count = TRUE
)
# $n
# [1] 70
```
<div id="workflow-ctis-example"></div>
* Queries to CTIS
Queries in the CTIS search interface (https://euclinicaltrials.eu/ctis-public/search) can be automatically copied to the clipboard so that a user can paste them into `queryterm`, see [here](#2-script-to-automatically-copy-users-query-from-web-browser). Subsequent to the relaunch of CTIS on 2024-07-24, there are now more than 11,050 trials publicly accessible in CTIS. See [below](#documents-example) for how to download documents from CTIS.
```{r load_ctis}
# See how many trials are in CTIS publicly accessible:
ctrLoadQueryIntoDb(
queryterm = "",
register = "CTIS",
only.count = TRUE
)
# $n
# [1] 11057
# Trials in therapeutic area neoplasms (ICD C04):
ctrLoadQueryIntoDb(
queryterm = 'searchCriteria={"therapeuticAreaCode":[4]}',
register = "CTIS",
only.count = TRUE
)
# * Found search query from CTIS: searchCriteria={"therapeuticAreaCode":[4]}
# * Checking trials in CTIS, found 3532 trials
# = Not done (only.count = TRUE): Imported 3532 trial(s)
# $n
# [1] 3622
```
<div id="workflow-data-model"></div>
* Inspect and search structure of trial information
For a given trial, function [ctrShowOneTrial()](https://rfhb.github.io/ctrdata/reference/ctrShowOneTrial.html) enables the user to visualise the hiearchy of fields and contents in the user's local web browser, to search for field names and field values, and to select and copy selected fields' names for use with function [dbGetFieldsIntoDf()](https://rfhb.github.io/ctrdata/reference/dbGetFieldsIntoDf.html).
```{r inspect_trial}
# This opens a local browser for user interaction.
# If the trial identifier (_id) is not found in the specified
# collection, it will be retrieved from the relevant register.
ctrShowOneTrial(
identifier = "2024-518931-12-00",
con = db
)
```
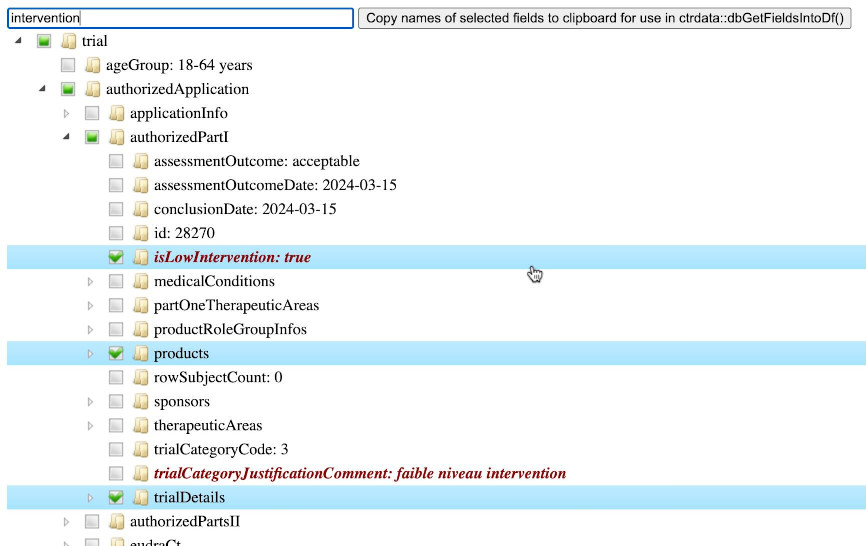
<div id="workflow-across-registers-example"></div>
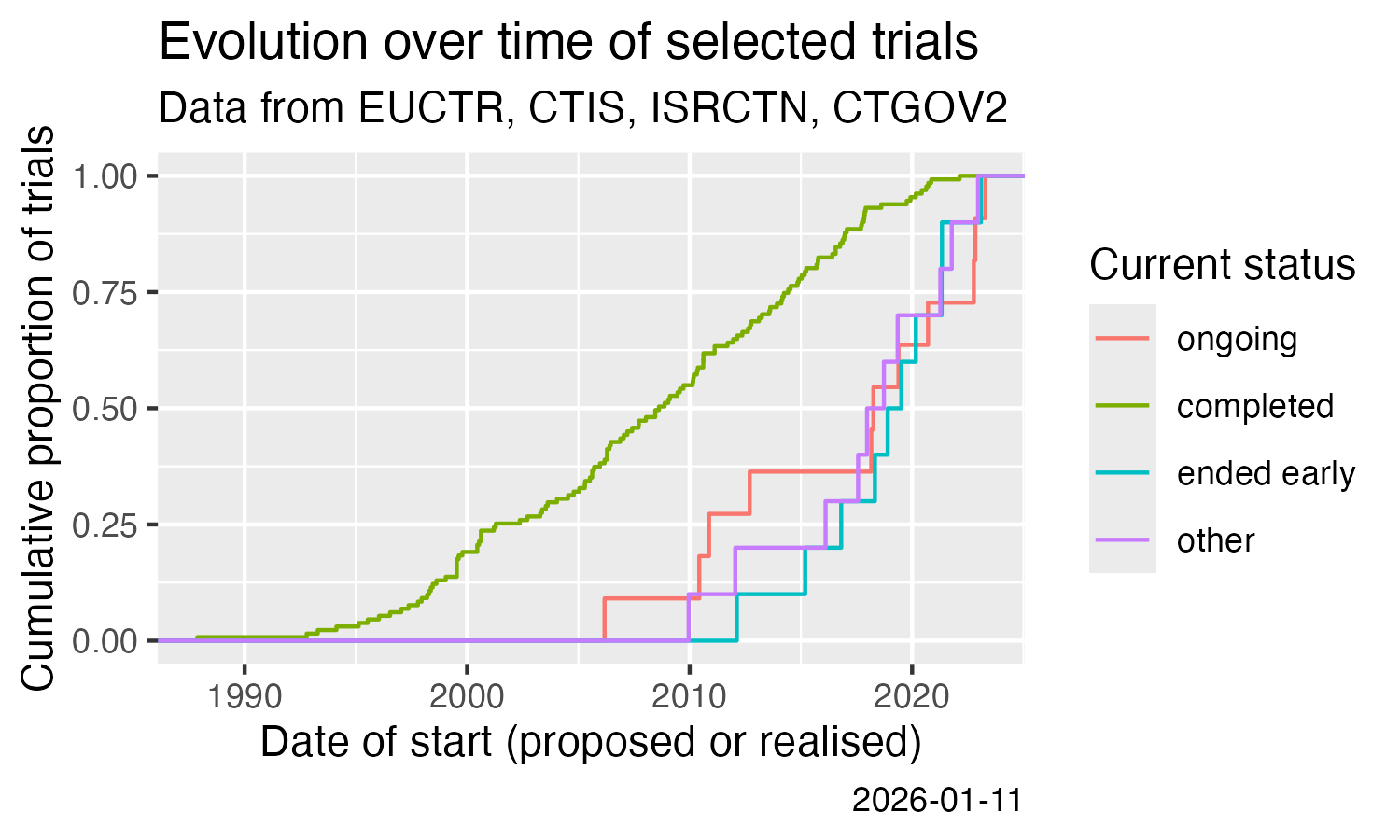
* Analysis across registers
Show cumulative start of trials over time. This uses the calculation of one of the trial concepts that are implemented in `ctrdata` since version 1.21.0.
```{r analyse_across_trials}
# explore 20 pre-defined concepts for
# trial analysis across registers
help("ctrdata-trial-concepts")
# use helper packages
library(dplyr)
library(ggplot2)
# calculate concept across registers
result <- dbGetFieldsIntoDf(
calculate = c(
"f.statusRecruitment",
"f.startDate"),
con = db
)
# not using trial concept f.isUniqueTrial
# but using dbFindIdsUniqueTrials() in
# order to prefer CTIS over EUCTR
result$.isUniqueTrial <- result$`_id` %in%
dbFindIdsUniqueTrials(
preferregister = c("CTGOV2", "CTIS", "EUCTR", "CTGOV", "ISRCTN"),
con = db
)
# plot concepts
result %>%
filter(.isUniqueTrial) %>%
ggplot() +
stat_ecdf(aes(
x = .startDate,
colour = .statusRecruitment
)) +
labs(
title = "Evolution over time of selected trials",
subtitle = "Data from EUCTR, CTIS, ISRCTN, CTGOV2",
x = "Date of start (proposed or realised)",
y = "Cumulative proportion of trials",
colour = "Current status",
caption = Sys.Date()
)
ggsave(
filename = "man/figures/README-ctrdata_across_registers.png",
width = 5, height = 3, units = "in"
)
```
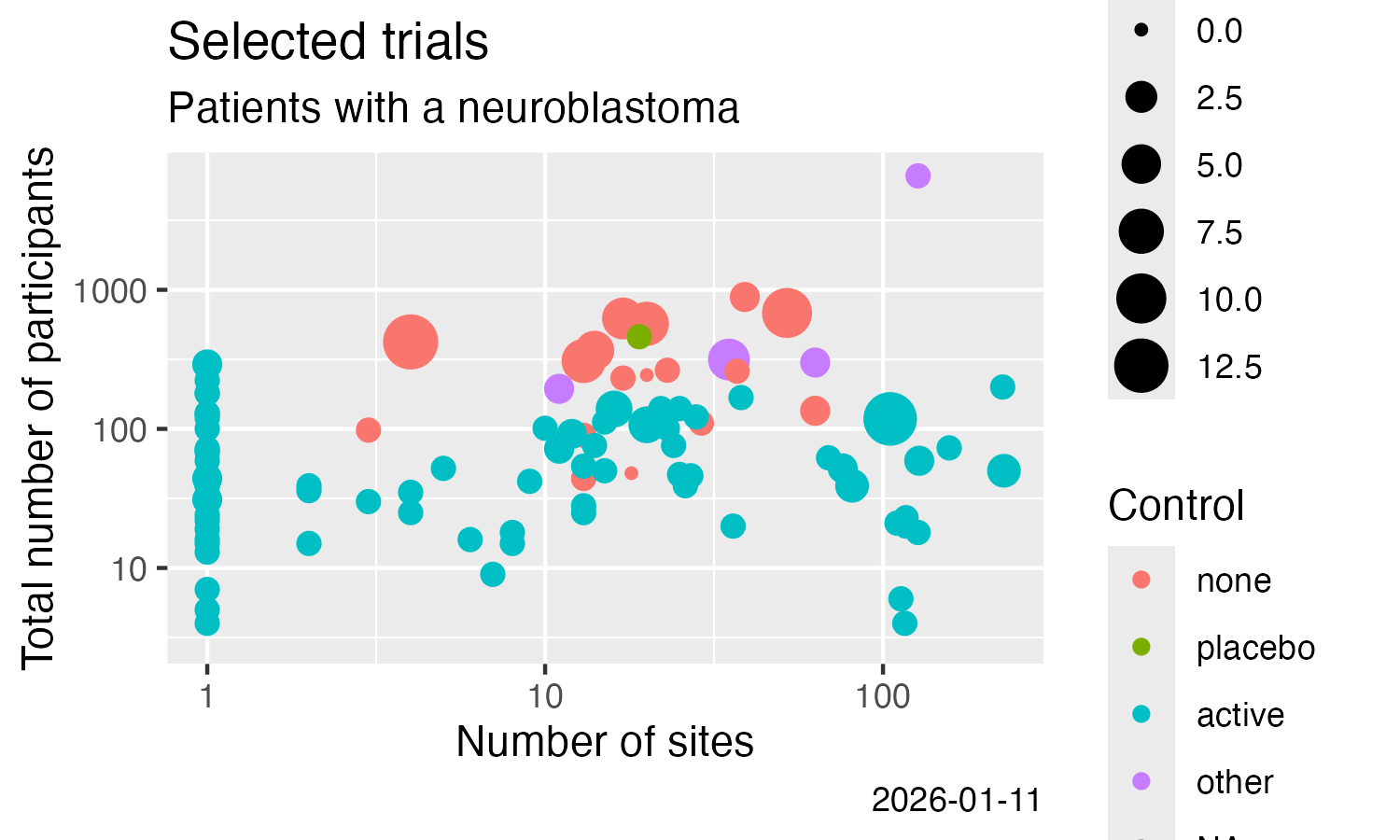
* Result-related trial information
Analyse some simple result details, here from CTGOV2 (see this [vignette](https://rfhb.github.io/ctrdata/articles/ctrdata_summarise.html) for more examples):
```{r analyse_results}
# use helper packages
library(dplyr)
library(ggplot2)
# calculate concept across registers
result <- dbGetFieldsIntoDf(
calculate = c(
"f.numSites",
"f.sampleSize",
"f.controlType",
"f.isUniqueTrial",
"f.numTestArmsSubstances"
),
con = db
)
# To review trial concepts details, call 'help("ctrdata-trial-concepts")'
# Querying database (50 fields)...
# plot concept
result %>%
filter(.isUniqueTrial) %>%
ggplot() +
labs(
title = "Selected trials",
subtitle = "Patients with a neuroblastoma"
) +
geom_point(
mapping = aes(
x = .numSites,
y = .sampleSize,
size = .numTestArmsSubstances,
colour = .controlType
)
) +
scale_x_log10() +
scale_y_log10() +
labs(
x = "Number of sites",
y = "Total number of participants",
colour = "Control",
size = "# Treatments",
caption = Sys.Date()
)
ggsave(
filename = "man/figures/README-ctrdata_results_neuroblastoma.png",
width = 5, height = 3, units = "in"
)
```
<div id="documents-example"></div>
* Download documents: retrieve protocols, statistical analysis plans and other documents into the local folder `./files-.../`
```{r load_documents}
### EUCTR document files can be downloaded when results are requested
# All files are downloaded and saved (documents.regexp is not used with EUCTR)
ctrLoadQueryIntoDb(
queryterm = "query=cancer&age=under-18&phase=phase-one",
register = "EUCTR",
euctrresults = TRUE,
euctrprotocolsall = FALSE, # new since 2025-07-20, loads single
# instead of all available country versions of a trial in EUCTR
documents.path = "./files-euctr/",
con = db
)
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one
# * Checking trials in EUCTR, found 249 trials
# - Running with euctrresults = TRUE to download documents
# - Created directory ./files-euctr/
# - Downloading in 13 batch(es) (20 trials each; estimate: 30 MB)
# - Downloading 249 records of 249 trials (estimate: 10 s)
# - Converting to NDJSON (estimate: 0.7 s)...
# - Importing records into database...
# = Imported or updated 249 records on 249 trial(s)
# * Checking results if available from EUCTR for 249 trials:
# - Downloading results...
# - Downloading documents into 'documents.path' = ./files-euctr
# - Extracting results (. = data, F = file[s] and data, x = none): F . . . . F . . F F . F . . . . F . . . . F . . . . . . . F . F . . F . . . . . . . . F F . . . . . . . . . . . . . . . . . . . F F . F . . . . . . . . . . . . . . . . . . . . . . . . . . F . . . . . . . . . . F F . . . . . . . . . . . . . . F . . . . F . . F . . . F . . . .
# - Data found for 138 trials
# - Converting to NDJSON (estimate: 4 s)...
# - Importing 138 results into database (may take some time)...
# - Results history: not retrieved (euctrresultshistory = FALSE)
# = Imported or updated results for 138 trials
# = Documents saved in './files-euctr'
# No history found in expected format.
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 249
### CTGOV files are downloaded, here corresponding to the default of
# documents.regexp = "prot|sample|statist|sap_|p1ar|p2ars|icf|ctalett|lay|^[0-9]+ "
ctrLoadQueryIntoDb(
queryterm = "cond=Neuroblastoma&type=Intr&recrs=e&phase=1&u_prot=Y&u_sap=Y&u_icf=Y",
register = "CTGOV",
documents.path = "./files-ctgov/",
con = db
)
# Since 2024-06-25, the classic CTGOV servers are no longer available. Package ctrdata has translated the classic CTGOV query URL from this call of function ctrLoadQueryIntoDb(queryterm = ...) into a query URL that works with the current CTGOV2. This is printed below and is also part of the return value of this function, ctrLoadQueryIntoDb(...)$url. This URL can be used with ctrdata functions. Note that the fields and data schema of trials differ between CTGOV and CTGOV2.
#
# Replace this URL:
#
# https://classic.clinicaltrials.gov/ct2/results?cond=Neuroblastoma&type=Intr&recrs=e&phase=1&u_prot=Y&u_sap=Y&u_icf=Y
#
# with this URL:
#
# https://clinicaltrials.gov/search?cond=Neuroblastoma&aggFilters=phase:2,docs:prot sap icf,studyType:int,status:com
#
# * Found search query from CTGOV2: cond=Neuroblastoma&aggFilters=phase:2,docs:prot sap icf,studyType:int,status:com
# * Checking trials in CTGOV2, found 35 trials
# - Downloading in 1 batch(es) (max. 1000 trials each; estimate: 3.5 MB total)
# - Load and convert batch 1...
# - Importing records into database...
# JSON file #: 1 / 1
# * Checking for documents...
# - Getting links to documents
# - Downloading documents into 'documents.path' = ./files-ctgov/
# - Created directory ./files-ctgov
# - Applying 'documents.regexp' to 45 missing documents
# - Creating subfolder for each trial
# = Newly saved 45 document(s) for 35 trial(s); 0 of such document(s) for 0 trial(s) already existed in ./files-ctgov
# = Imported or updated 35 trial(s)
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 35
### CTGOV2 files are downloaded, using the default of documents.regexp
ctrLoadQueryIntoDb(
queryterm = "https://clinicaltrials.gov/search?cond=neuroblastoma&aggFilters=phase:1,results:with",
documents.path = "./files-ctgov2/",
con = db
)
# * Found search query from CTGOV2: cond=neuroblastoma&aggFilters=phase:1,results:with
# * Checking trials in CTGOV2, found 42 trials
# - Downloading in 1 batch(es) (max. 1000 trials each; estimate: 4.2 MB total)
# - Load and convert batch 1...
# - Importing records into database...
# JSON file #: 1 / 1
# * Checking for documents...
# - Getting links to documents
# - Downloading documents into 'documents.path' = ./files-ctgov2/
# - Created directory ./files-ctgov2
# - Applying 'documents.regexp' to 46 missing documents
# - Creating subfolder for each trial
# = Newly saved 46 document(s) for 29 trial(s); 0 of such document(s) for 0 trial(s) already existed in ./files-ctgov2
# = Imported or updated 42 trial(s)
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 42
### ISRCTN files are downloaded, using the default of documents.regexp
ctrLoadQueryIntoDb(
queryterm = "https://www.isrctn.com/search?q=alzheimer",
documents.path = "./files-isrctn/",
con = db
)
# * Found search query from ISRCTN: q=alzheimer
# * Checking trials in ISRCTN, found 350 trials
# - Downloading trial file (estimate: 6 MB)...
# - Converting to NDJSON (estimate: 2 s)...
# - Importing records into database...
# * Checking for documents...
# - Getting links to documents from data . correct with web pages . . . . . . . .
# - Downloading documents into 'documents.path' = ./files-isrctn/
# - Created directory ./files-isrctn
# - Applying 'documents.regexp' to 61 missing documents
# - Creating subfolder for each trial
# = Newly saved 37 document(s) for 16 trial(s); 0 of such document(s) for 0 trial(s) already existed in ./files-isrctn
# = Imported or updated 350 trial(s)
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 350
### CTIS files are downloaded, using a specific documents.regexp
ctrLoadQueryIntoDb(
queryterm = paste0(
"https://euclinicaltrials.eu/ctis-public/search#",
'searchCriteria={"containAny":"cancer","status":[8]}'),
documents.path = "./files-ctis/",
documents.regexp = "^Prtcl-Extrct",
con = db
)
# * Found search query from CTIS: searchCriteria={"containAny":"cancer","status":[8]}
# * Checking trials in CTIS, found 440 trials
# - Downloading and processing trial list... (estimate: 0.8 s)
# - Downloading and processing trial data... (estimate: 60 MB)
# - Importing records into database...
# - Updating with additional data: .
# * Checking for documents . . . . .
# - Downloading documents into 'documents.path' = ./files-ctis/
# - Created directory ./files-ctis
# - Applying 'documents.regexp' to 8125 missing documents
# - Creating subfolder for each trial
# - Downloading 2 missing documents . .
# = Newly saved 2 document(s) for 2 trial(s); 0 of such document(s) for 0 trial(s) already existed in ./files-ctis
# = Imported 440, updated 440 record(s) on 440 trial(s)
# Updated history ("meta-info" in "collection_name")
# $n
# [1] 440
```
```{r cleanup, include=FALSE}
# cleanup
unlink("database_name.sql")
unlink("./files-ctis/", recursive = TRUE)
unlink("./files-euctr/", recursive = TRUE)
unlink("./files-ctgov/", recursive = TRUE)
unlink("./files-ctgov2/", recursive = TRUE)
unlink("./files-isrctn/", recursive = TRUE)
```
## Tests and coverage
See also [https://app.codecov.io/gh/rfhb/ctrdata/tree/master/R](https://app.codecov.io/gh/rfhb/ctrdata/tree/master/R)
```{r}
# 2026-03-07
tinytest::test_all()
# test_ctrdata_duckdb_ctgov2.R.. 79 tests OK 55.3s
# test_ctrdata_function_activesubstance.R 4 tests OK 0.8s
# test_ctrdata_function_ctrgeneratequeries.R 10 tests OK 0.2s
# test_ctrdata_function_params.R 25 tests OK 1.5s
# test_ctrdata_function_trial-concepts.R 93 tests OK 1.7s
# test_ctrdata_function_various.R 79 tests OK 13.4s
# test_ctrdata_mongo_local_euctr.R 117 tests OK 44.9s
# test_ctrdata_mongo_remote_ro.R 4 tests OK 6.0s
# test_ctrdata_sqlite_ctgov.R... 48 tests OK 30.8s
# test_ctrdata_sqlite_ctis.R.... 95 tests OK 7.4s
# test_ctrdata_sqlite_isrctn.R.. 47 tests OK 13.3s
# test_euctr_error_sample.R..... 8 tests OK 0.1s
# All ok, 609 results (10m 9.4s)
```
```{r}
# 2026-03-07
covr::package_coverage(path = ".", type = "tests")
# ctrdata Coverage: 94.26%
# R/zzz.R: 55.56%
# R/ctrShowOneTrial.R: 84.21%
# R/ctrLoadQueryIntoDbEuctr.R: 87.97%
# R/ctrFindActiveSubstanceSynonyms.R: 88.89%
# R/ctrGetQueryUrl.R: 89.18%
# R/f_primaryEndpointResults.R: 89.24%
# R/dbGetFieldsIntoDf.R: 89.47%
# R/util_functions.R: 89.83%
# R/ctrLoadQueryIntoDbCtis.R: 90.00%
# R/f_sponsorType.R: 90.00%
# R/ctrRerunQuery.R: 92.22%
# R/ctrLoadQueryIntoDbCtgov2.R: 92.48%
# R/ctrLoadQueryIntoDbIsrctn.R: 95.68%
# R/dbFindFields.R: 95.88%
# R/ctrLoadQueryIntoDb.R: 96.40%
# R/dfMergeVariablesRelevel.R: 96.55%
# R/ctrGenerateQueries.R: 97.16%
# R/f_externalLinks.R: 97.30%
# R/ctrOpenSearchPagesInBrowser.R: 97.50%
# R/f_likelyPlatformTrial.R: 98.78%
# R/dbFindIdsUniqueTrials.R: 98.81%
# R/f_numTestArmsSubstances.R: 98.95%
# R/dbQueryHistory.R: 100.00%
# R/dfName2Value.R: 100.00%
# R/dfTrials2Long.R: 100.00%
# R/f_assignmentType.R: 100.00%
# R/f_controlType.R: 100.00%
# R/f_hasResults.R: 100.00%
# R/f_isMedIntervTrial.R: 100.00%
# R/f_isUniqueTrial.R: 100.00%
# R/f_numSites.R: 100.00%
# R/f_primaryEndpointDescription.R: 100.00%
# R/f_resultsDate.R: 100.00%
# R/f_sampleSize.R: 100.00%
# R/f_startDate.R: 100.00%
# R/f_statusRecruitment.R: 100.00%
# R/f_trialObjectives.R: 100.00%
# R/f_trialPhase.R: 100.00%
# R/f_trialPopulation.R: 100.00%
# R/f_trialTitle.R: 100.00%
```
## Future features
* See project outline https://github.com/users/rfhb/projects/1
* Authentication, expected to be required by CTGOV2; specifications not yet known (work not yet started).
* Explore further registers (exploration is continually ongoing; added value, terms and conditions for programmatic access vary; no clear roadmap is established yet).
## Acknowledgements
* Data providers and curators of the clinical trial registers. Please review and respect their copyrights and terms and conditions, see `ctrOpenSearchPagesInBrowser(copyright = TRUE)`.
* Package `ctrdata` has been made possible building on the work done for
[R](https://www.r-project.org/),
[dplyr](https://cran.r-project.org/package=dplyr),
[duckdb](https://cran.r-project.org/package=duckdb),
[htmlwidgets](https://cran.r-project.org/package=htmlwidgets),
[httr2](https://cran.r-project.org/package=httr2),
[jqr](https://cran.r-project.org/package=jqr),
[jsonlite](https://cran.r-project.org/package=jsonlite),
[lubridate](https://cran.r-project.org/package=lubridate),
[mongolite](https://cran.r-project.org/package=mongolite),
[nodbi](https://cran.r-project.org/package=nodbi),
[readr](https://cran.r-project.org/package=readr).
[rlang](https://cran.r-project.org/package=rlang),
[RPostgres](https://cran.r-project.org/package=RPostgres),
[RSQLite](https://CRAN.R-project.org/package=RSQLite),
[stringdist](https://cran.r-project.org/package=stringdist) and
[stringi](https://cran.r-project.org/package=stringi) and
[tidyr](https://cran.r-project.org/package=tidyr),
[V8](https://cran.r-project.org/package=V8),
[xml2](https://cran.r-project.org/package=xml2).
## Issues and notes
* Information in trial registers may not be fully correct; see for example [this publication on CTGOV](https://doi.org/10.1136/bmj.k1452).
* A warning may be issued and a record not imported if the complexity of the XML content is too high for processing. The issue can be resolved by increasing in the operating system the stack size available to R, see: https://github.com/rfhb/ctrdata/issues/22
* Please file issues and bugs [here](https://github.com/rfhb/ctrdata/issues).
* Check out any relevant closed issues, e.g. on [C stack usage too close to the limit](https://github.com/rfhb/ctrdata/issues/22) and on a [SSL certificate problem: unable to get local issuer certificate](https://github.com/rfhb/ctrdata/issues/19#issuecomment-820127139).