Signalling networks are formed as a result of interacting cell signalling pathways. One important example of a cell signalling network is that formed by kinase proteins. These enzymes interact with other kinases or with themselves by phosphorylating hydroxyl groups of some residues. Given that there are approximately 518 human kinases, by controlling the activity and function of various proteins, they play a key role in the regulation of overall cellular function. Therefore, disruption of kinase signalling networks affects the cell physiology and has been known to have implications in cancer and neurodegenerative diseases.
KiNet constructs and visualises kinase signalling network using a dataset of pre-identified proteins and extracted UniProt PTM data. The simple-interaction-format (.SIF) file produced by the script is suitable as input into Cytoscape, a network generator application.
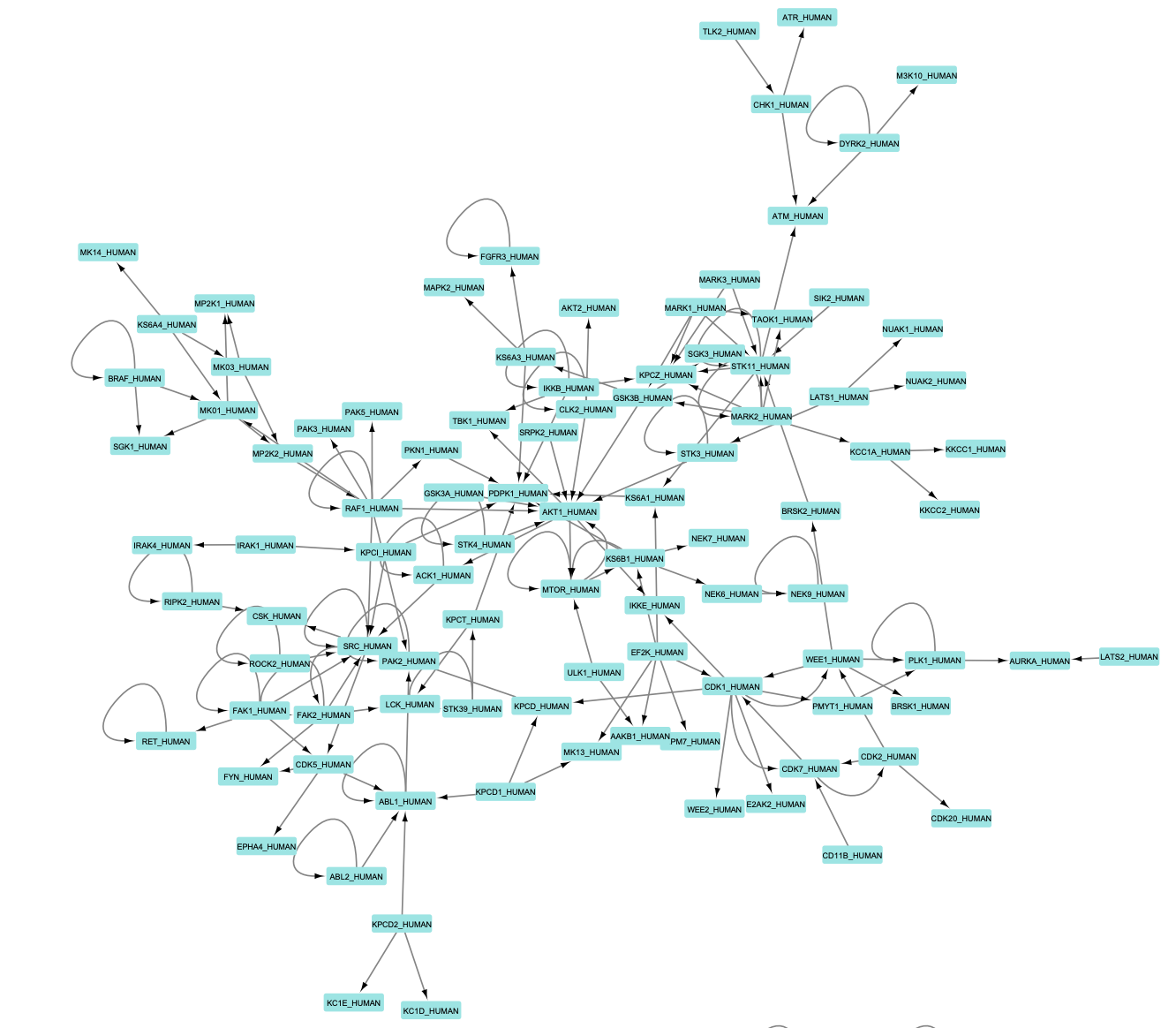
A graphical representation of the .SIF file generated using the project's example dataset is shown below:
- Python 2