+__date__ = "10 Nov 2018"
+__author__ = "Baris Ozmen"
+
+
+"""Collects plotting functions to be used
+"""
+
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+import altair as alt
+
+import sys
+
+sys.path.append("../")
+
+from lib.decorators import Reporter
+
+timer = Reporter.timer
+counter = Reporter.counter
+
+plt.interactive(False)
+
+
+[docs]class PlotOp:
+
"""Plotting Operations class. Keeps static plot functions.
+
+
Each function is conforming as much as possible to "Functional Programming Standards" (https://en.wikipedia.org/wiki/Functional_programming).
+
Each function, hence, do not affected by any variable not given to it as argument (with exception of influences from
+
appealed external libraries, such as matplotlib, altair, and bokeh) .
+
"""
+
+
[docs] @staticmethod
+
@timer
+
@counter
+
def plot_heatmap(
+
value_matrix,
+
xlabels,
+
ylabels,
+
min_value=-1,
+
max_value=1,
+
labels_fontsize=24,
+
xlabels_fontsize=None,
+
ylabels_fontsize=None,
+
colorbar_labels_fontsize=None,
+
sg_length=None,
+
figsize=None,
+
save_fig_to=None,
+
):
+
+
"""Plots heatmap of a given 2d array
+
+
Args:
+
value_matrix (numpy.array): matrix to plot its heatmap
+
xlabels (list):
+
ylabels (list):
+
min_value (float):
+
max_value (float):
+
labels_fontsize (int): determines fontsize of text in the plot.
+
+
xlabels_fontsize (int): determines fontsize of x axis labels. If not given,
+
value of `labels_fontsize` argument is used.
+
+
ylabels_fontsize (int): determines fontsize of y axis labels. If not given,
+
value of `labels_fontsize` argument is used.
+
+
colorbar_labels_fontsize (int): determines fontsize of colorbar labels. If
+
not given, value of `labels_fontsize` argument is used.
+
+
sg_length (int): slopper over grid line lenght. If not given, it is
+
taken as 10% of length of x or y axes whichever is smaller.
+
+
figsize (tuple): determines size of the plot
+
"""
+
+
if xlabels_fontsize is None:
+
xlabels_fontsize = labels_fontsize
+
+
if ylabels_fontsize is None:
+
ylabels_fontsize = labels_fontsize
+
+
if colorbar_labels_fontsize is None:
+
colorbar_labels_fontsize = labels_fontsize - 2
+
+
if figsize is None:
+
figsize = (int(len(xlabels) / 2), int(len(ylabels) / 2))
+
print("figsize is {}".format(figsize))
+
+
fig = plt.figure(figsize=figsize)
+
+
################
+
# Plot heatmap #
+
################
+
im = plt.imshow(
+
value_matrix,
+
cmap="bwr",
+
interpolation="nearest",
+
vmin=min_value,
+
vmax=max_value,
+
)
+
+
ax = plt.gca()
+
+
ax.xaxis.set_ticks_position("top")
+
ax.yaxis.set_ticks_position("left")
+
ax.set_xticks(range(len(xlabels)))
+
ax.set_xticklabels(xlabels, rotation=90, fontsize=xlabels_fontsize)
+
ax.set_yticks(range(len(ylabels)))
+
ax.set_yticklabels(ylabels, rotation=0, fontsize=ylabels_fontsize)
+
+
ax.spines["left"].set_visible(False)
+
ax.spines["bottom"].set_visible(False)
+
+
xlim = ax.get_xlim()
+
ylim = ax.get_ylim()
+
+
xlength = xlim[1] - xlim[0]
+
ylength = ylim[1] - ylim[0]
+
+
if sg_length is None:
+
sg_length = min(xlength, ylength) * 0.1
+
+
x_micro_grid = np.arange(0, len(xlabels) + 1, 1) + 0.5
+
y_micro_grid = np.arange(0, len(ylabels) + 1, 1) + 0.5
+
+
x_macro_grid = np.arange(0, len(xlabels) + 1, 4) - 0.5
+
y_macro_grid = np.arange(0, len(ylabels) + 1, 4) - 0.5
+
+
for x in x_micro_grid:
+
plt.plot([x, x], ylim, "k-", lw=0.5)
+
+
for y in y_micro_grid:
+
plt.plot(xlim, [y, y], "k-", lw=0.5)
+
+
for x in x_macro_grid:
+
lines = plt.plot([x, x], [ylim[0], ylim[1] + sg_length], "k-", lw=2)
+
lines[0].set_clip_on(False)
+
+
for y in y_macro_grid:
+
lines = plt.plot([xlim[0] + sg_length, xlim[1]], [y, y], "k-", lw=2)
+
lines[0].set_clip_on(False)
+
+
plt.xlim(xlim)
+
plt.ylim(ylim)
+
+
colorbar_axes = fig.add_axes([1, 0.13, 0.02, 0.8])
+
+
colorbar_axes.tick_params(labelsize=labels_fontsize - 2)
+
+
plt.colorbar(im, cax=colorbar_axes)
+
+
if save_fig_to is not None:
+
plt.savefig(save_fig_to)
+
+
plt.show()
+
+
[docs] @staticmethod
+
@timer
+
@counter
+
def plot_heatmap_of_uppertriangle(
+
value_matrix,
+
xlabels,
+
ylabels,
+
min_value=-1,
+
max_value=1,
+
labels_fontsize=18,
+
xlabels_fontsize=None,
+
ylabels_fontsize=None,
+
colorbar_labels_fontsize=None,
+
sg_length=None,
+
figsize=None,
+
save_fig_to=None,
+
):
+
+
"""Plots heatmap of a given 2d array
+
+
Args:
+
value_matrix (numpy.array): matrix to plot its heatmap
+
xlabels (list):
+
ylabels (list):
+
min_value (float):
+
max_value (float):
+
labels_fontsize (int): determines fontsize of text in the plot.
+
+
xlabels_fontsize (int): determines fontsize of x axis labels. If not given,
+
value of `labels_fontsize` argument is used.
+
+
ylabels_fontsize (int): determines fontsize of y axis labels. If not given,
+
value of `labels_fontsize` argument is used.
+
+
colorbar_labels_fontsize (int): determines fontsize of colorbar labels. If
+
not given, value of `labels_fontsize` argument is used.
+
+
sg_length (int): slopper over grid line lenght. If not given, it is
+
taken as 10% of length of x or y axes whichever is smaller.
+
+
figsize (tuple): determines size of the plot
+
"""
+
+
if xlabels_fontsize is None:
+
xlabels_fontsize = labels_fontsize
+
+
if ylabels_fontsize is None:
+
ylabels_fontsize = labels_fontsize
+
+
if colorbar_labels_fontsize is None:
+
colorbar_labels_fontsize = labels_fontsize - 2
+
+
if figsize is None:
+
figsize = (int(len(xlabels) / 2), int(len(ylabels) / 2))
+
print("figsize is {}".format(figsize))
+
+
# get upper-triangle
+
value_matrix = np.triu(value_matrix, k=1)
+
+
fig = plt.figure(figsize=figsize)
+
+
################
+
# Plot heatmap #
+
################
+
im = plt.imshow(
+
value_matrix,
+
cmap="bwr",
+
interpolation="nearest",
+
vmin=min_value,
+
vmax=max_value,
+
)
+
+
ax = plt.gca()
+
+
ax.xaxis.set_ticks_position("top")
+
ax.yaxis.set_ticks_position("right")
+
ax.set_xticks(range(len(xlabels)))
+
ax.set_xticklabels(xlabels, rotation=90, fontsize=xlabels_fontsize)
+
ax.set_yticks(range(len(ylabels)))
+
ax.set_yticklabels(ylabels, rotation=0, fontsize=ylabels_fontsize)
+
+
ax.spines["left"].set_visible(False)
+
ax.spines["bottom"].set_visible(False)
+
+
xlim = ax.get_xlim()
+
ylim = ax.get_ylim()
+
+
xlength = xlim[1] - xlim[0]
+
ylength = ylim[1] - ylim[0]
+
+
if sg_length is None:
+
sg_length = min(xlength, ylength) * 0.1
+
+
x_micro_grid = np.arange(0, len(xlabels) + 1, 1) + 0.5
+
y_micro_grid = np.arange(0, len(ylabels) + 1, 1) + 0.5
+
+
x_macro_grid = np.arange(0, len(xlabels) + 1, 4) - 0.5
+
y_macro_grid = np.arange(0, len(ylabels) + 1, 4) - 0.5
+
+
for x in x_micro_grid:
+
plt.plot([x, x], [x, ylim[1]], "k-", lw=0.5)
+
+
for y in y_micro_grid:
+
plt.plot([y, xlim[1]], [y, y], "k-", lw=0.5)
+
+
for x in x_macro_grid:
+
lines = plt.plot([x, x], [x, ylim[1] + sg_length], "k-", lw=2)
+
lines[0].set_clip_on(False)
+
+
for y in y_macro_grid:
+
lines = plt.plot([y, xlim[1] - sg_length], [y, y], "k-", lw=2)
+
lines[0].set_clip_on(False)
+
+
plt.xlim(xlim)
+
plt.ylim(ylim)
+
+
colorbar_axes = fig.add_axes([0, 0.13, 0.02, 0.6])
+
colorbar_axes.tick_params(labelsize=labels_fontsize - 2)
+
+
plt.colorbar(im, cax=colorbar_axes)
+
+
if save_fig_to is not None:
+
plt.savefig(save_fig_to + ".png")
+
+
plt.show()
+
+
[docs] @staticmethod
+
@timer
+
@counter
+
def plot_heatmap_with_altair(
+
df,
+
xcol="event1",
+
ycol="event2",
+
valuecol="phi",
+
minval=-1,
+
maxval=1,
+
tooltip=None,
+
savefigto=None,
+
):
+
"""
+
+
Args:
+
df (pandas.DataFrame): dataframe to be drawn
+
xcol (str): `df` column for x-axis
+
ycol (str): `df` column for
+
valuecol (str): `df` column for value (colored)
+
tooltip (list): `df` columns to be shown when blocks are hovered by mouse
+
savefigto (str): address that figure will be saved
+
"""
+
# Argument operations
+
assert xcol in df.columns
+
assert ycol in df.columns
+
assert savefigto is not None
+
+
if tooltip is None:
+
tooltip = [xcol, ycol]
+
assert all(item in df.columns for item in tooltip)
+
+
import altair as alt
+
+
color_scheme = "redblue"
+
+
selection_by_x = alt.selection_multi(fields=[xcol], nearest=False)
+
+
selection_by_y = alt.selection_multi(fields=[ycol], nearest=False)
+
+
heatmap_base = (
+
alt.Chart(df)
+
.mark_rect()
+
.encode(
+
x=xcol,
+
y=ycol,
+
color=alt.Color(
+
"{}:Q".format(valuecol),
+
scale=alt.Scale(scheme=color_scheme, domain=[maxval, minval]),
+
),
+
tooltip=tooltip,
+
)
+
)
+
+
heatmap1 = heatmap_base.encode(
+
opacity=alt.condition(selection_by_x, alt.value(1), alt.value(0.2))
+
).add_selection(selection_by_x)
+
+
heatmap2 = heatmap_base.encode(
+
opacity=alt.condition(selection_by_y, alt.value(1), alt.value(0.2))
+
).add_selection(selection_by_y)
+
+
whole_chart = alt.layer(heatmap1, heatmap2).configure_view(strokeWidth=0)
+
+
# whole_chart.save(savefigto + '.png')
+
+
whole_chart.save(savefigto + ".html")
+
+
return whole_chart
+
+
[docs] @staticmethod
+
@timer
+
@counter
+
def plot_mutation_percentage(df):
+
"""Plots percentage of patients having misssense or nonsense mutation, as a vertical barplot.
+
+
Args:
+
dataframe:
+
columns: hgnc_symbol, percentage_missense, percentage_nonsense
+
"""
+
+
# data to plot
+
n_groups = df["hgnc_symbol"].unique().__len__()
+
mis_percentages = df["percentage_missense"].values
+
non_percentages = df["percentage_nonsense"].values
+
+
# create plot
+
fig, ax = plt.subplots(figsize=(5, 30))
+
+
index = np.arange(n_groups)
+
bar_width = 0.25
+
opacity = 1
+
+
rects1 = plt.barh(
+
index,
+
non_percentages,
+
bar_width,
+
alpha=opacity,
+
color="salmon",
+
label="Nonsense",
+
zorder=2,
+
)
+
+
rects2 = plt.barh(
+
index + bar_width,
+
mis_percentages,
+
bar_width,
+
alpha=opacity,
+
color="c",
+
label="Missense",
+
zorder=2,
+
)
+
+
plt.title(
+
"LUAD Driver Genes by Proportion of Samples Mutated \n (ordered alphabetically)"
+
)
+
+
plt.xticks(np.arange(0, 35, 5), [str(i) + "%" for i in np.arange(0, 35, 5)])
+
plt.yticks(index + bar_width / 2, df["hgnc_symbol"].unique())
+
ax.set_xticks(np.arange(0, 35, 1), minor=True)
+
plt.legend()
+
+
# ax.yaxis.grid()
+
ax.xaxis.grid(True, "minor", linestyle="--", lw=0.2, color="black", zorder=0)
+
ax.xaxis.grid(True, "major", linestyle="-", lw=0.3, color="black", zorder=0)
+
+
x0, x1 = plt.xlim()
+
y0, y1 = plt.ylim()
+
+
for ylevel in np.arange(5, y1, 10):
+
+
plt.plot(
+
[x0, x1 + 1],
+
[ylevel + 0.5, ylevel + 0.5],
+
color="black",
+
alpha=0.3,
+
lw=1.2,
+
ls="--",
+
)
+
+
for i in np.arange(5, 35, 5):
+
plt.text(i - 0.8, ylevel + 0.6, str(i) + "%", alpha=0.4)
+
+
plt.xlim((x0, x1 - 1))
+
+
plt.ylim(y0 + 2, y1 - 1.5)
+
+
plt.tight_layout()
+
+
[docs] @staticmethod
+
@timer
+
@counter
+
def plot_volcano(
+
df,
+
xcol,
+
ycol,
+
xcolrange=[-1, 1],
+
xtitle=None,
+
ytitle=None,
+
ythreshold=2,
+
clampthreshold=400,
+
tooltip=[],
+
topk=5,
+
figsize=(600, 600),
+
titlefontsize=18,
+
titlefontweight="normal",
+
labelfontsize=14,
+
savefigto="volcano_plot",
+
show=False,
+
):
+
"""Plots volcano figure and saves it to given address
+
+
Args:
+
df (pandas.DataFrame): data
+
xcol (str): column for x-axis
+
ycol (str):
+
xcolrange (tuple):
+
xtitle (str):
+
ytitle (str):
+
ythreshold int:
+
tooltip (list):
+
topk int:
+
figsize (tuple):
+
titlefontsize (int):
+
titlefontweight (int or str):
+
labelfontsize (int):
+
savefigto (str):
+
"""
+
assert xcol in df.columns
+
assert ycol in df.columns
+
assert all(item in df.columns for item in tooltip)
+
+
# CLAMP fisher-exact values
+
if np.sum(df[ycol] > clampthreshold) > 0:
+
df["fisher_exact"] = np.where(
+
df["fisher_exact"] > clampthreshold, clampthreshold, df["fisher_exact"]
+
)
+
print(
+
"Datapoints' fisher-exact values clamped at {}!".format(clampthreshold)
+
)
+
+
xAxis = alt.Axis(title=xtitle)
+
yAxis = alt.Axis(title=ytitle)
+
+
X = alt.X(xcol, scale=alt.Scale(domain=xcolrange), axis=xAxis)
+
Y = alt.Y(ycol, scale=alt.Scale(domain=[0, df[ycol].max()]), axis=yAxis)
+
+
points_below = (
+
alt.Chart(df[df[ycol] < ythreshold])
+
.encode(x=X, y=Y, tooltip=tooltip)
+
.mark_circle(size=300, opacity=0.5, color="blue")
+
.interactive()
+
)
+
+
points_above = (
+
alt.Chart(df[df[ycol] >= ythreshold])
+
.encode(x=X, y=Y, tooltip=tooltip)
+
.mark_circle(size=300, opacity=0.5, color="red")
+
.interactive()
+
)
+
+
top_k_points = (
+
alt.Chart(df.nlargest(topk, ycol))
+
.encode(x=X, y=Y)
+
.mark_circle(size=0, opacity=0)
+
.mark_text(align="right", baseline="bottom", dx=35, dy=-17)
+
.encode(text="pairs")
+
)
+
+
rule1_data = pd.DataFrame([{"fisher_exact": ythreshold}])
+
rule1 = (
+
alt.Chart(rule1_data)
+
.mark_rule(color="black", opacity=1.0, strokeDash=[1, 1])
+
.encode(y=ycol)
+
)
+
+
rule2_data = pd.DataFrame([{xcol: 0}])
+
rule2 = (
+
alt.Chart(rule2_data).mark_rule(color="black", opacity=1.0).encode(x=xcol)
+
)
+
+
# Superimpose all charts
+
whole_chart = rule1 + rule2 + points_below + points_above + top_k_points
+
+
# adjust aesthetics
+
whole_chart = whole_chart.properties(
+
width=figsize[0], height=figsize[1]
+
).configure_axis(
+
titleFontSize=titlefontsize,
+
titleFontWeight=titlefontweight,
+
labelFontSize=labelfontsize,
+
)
+
+
# save as html
+
if savefigto is not None:
+
whole_chart.save(savefigto + ".html")
+
+
if show == True:
+
alt.renderers.enable("notebook")
+
return whole_chart
+
+
[docs] def pvalue_qq_plot(obs_pvalues, label_fontsize=20, ax=None, param_dict=None):
+
""" Draws a p-value QQ-plot given observed p-values
+
+
Args:
+
obs_pvalues (list): observed p-values
+
label_fontsize (int): fontsize of x and y axis labels
+
ax (matplotlib.axes._subplots.AxesSubplot): (optional) AxesSubplot object for plot to be drawn on.
+
"""
+
+
if ax is None:
+
fig = plt.figure(figsize=(6, 6))
+
ax = fig.gca()
+
+
obs_pvalues = np.array(obs_pvalues)
+
if obs_pvalues.ndim == 1:
+
xs = np.arange(1, len(obs_pvalues) + 1) / len(obs_pvalues)
+
ax.plot(-np.log10(xs), -np.log10(sorted(obs_pvalues)), ".", color="black")
+
else:
+
for pvalues in obs_pvalues:
+
print(len(pvalues))
+
xs = np.arange(1, len(pvalues) + 1) / len(pvalues)
+
ax.plot(-np.log10(xs), -np.log10(sorted(pvalues)), ".")
+
+
# see **kwargs parameter in matplotlib
+
# documentation (https://matplotlib.org/api/pyplot_api.html)
+
# for using param_dict more efficiently in the future
+
if "fillstyle" not in param_dict:
+
param_dict["fillstyle"] = "--"
+
if "color" not in param_dict:
+
param_dict["color"] = "gray"
+
+
ax.plot(-np.log10(xs), -np.log10(xs), **param_dict)
+
ax.set_xlabel("\n-log10 (Expected P-value)", fontsize=label_fontsize)
+
ax.set_ylabel("-log10 (Observed P-value)\n", fontsize=label_fontsize)
+
 +
+## Design goals
+DeepAugment is designed as a scalable and modular partner to AutoAugment ([Cubuk et al., 2018](https://arxiv.org/abs/1805.09501)). AutoAugment was one of the most exciting publications in 2018. It was the first method using Reinforcement Learning for this problem. AutoAugmentation, however, has no complete open-sourced implementation (controller module not available) preventing users to run it for their own datasets, and takes 15,000 iterations to learn (according to paper) augmentation policies, which requires massive computational resources. Thus most people could not benefit from it even if its source code would be fully available.
+
+DeepAugment addresses these two problems. Its main design goals are:
+1. **minimize the computational complexity of optimization while maintaining quality of results**
+2. **be modular and user-friendly**
+
+First goal is achieved by following changes compared to AutoAugment:
+1. Bayesian Optimization instead of Reinforcement Learning
+ * which requires much less number of iterations (~100 times)
+2. Minimized Child Model
+ * decreasing computational complexity of each training (~20 times)
+3. Less stochastic augmentation search space design
+ * decreasing number of iterations needed
+
+For achieving the second goal, user interface is designed in a way that it gives user broad configuration possibilities and model selections (e.g. selecting the child model or inputting a self-designed child model).
+
+## Importance
+### Practical importance
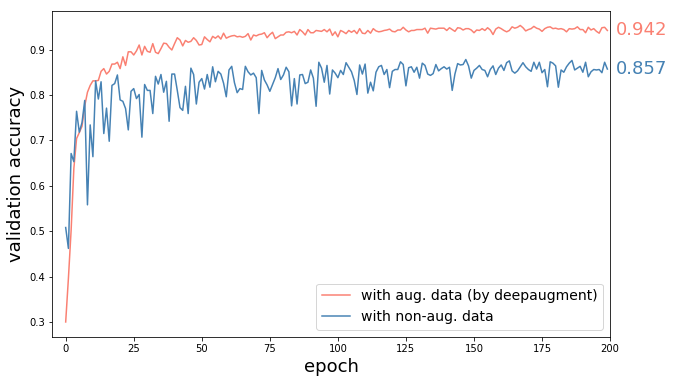
+DeepAugment makes optimization of data augmentation scalable, and thus enables users to optimize augmentation policies without needing massive computational resources.
+As an estimate of its computational cost, it takes **4.2 hours** (500 iterations) on CIFAR-10 dataset which costs around **$13** using AWS p3.x2large instance.
+### Academic importance
+To our knowledge, DeepAugment is the first method which utilizes Bayesian Optimization for the problem of data augmentation hyperparameter optimization.
+
+## How it works
+
+Three major components of DeepAugment are controller, augmenter, and child model. Overall workflow is that controller samples new augmentation policies, augmenter transforms images by the new policy, and child model is trained from scratch by augmented images. Then, a reward is calculated from child model's training history. This reward is returned back to the controller, and it updates its surrogate model with this reward and associated augmentation policy. Then, controller samples new policies again and same steps repeats. This process cycles until user-determined maximum number of iterations reached.
+
+Controller can be set for using either Bayesian Optimization (default) or Random Search. If set to Bayesian Optimization, samples new policies by a Random Forest Estimator and Expected Improvement acquisition function.
+
+
+
+## Design goals
+DeepAugment is designed as a scalable and modular partner to AutoAugment ([Cubuk et al., 2018](https://arxiv.org/abs/1805.09501)). AutoAugment was one of the most exciting publications in 2018. It was the first method using Reinforcement Learning for this problem. AutoAugmentation, however, has no complete open-sourced implementation (controller module not available) preventing users to run it for their own datasets, and takes 15,000 iterations to learn (according to paper) augmentation policies, which requires massive computational resources. Thus most people could not benefit from it even if its source code would be fully available.
+
+DeepAugment addresses these two problems. Its main design goals are:
+1. **minimize the computational complexity of optimization while maintaining quality of results**
+2. **be modular and user-friendly**
+
+First goal is achieved by following changes compared to AutoAugment:
+1. Bayesian Optimization instead of Reinforcement Learning
+ * which requires much less number of iterations (~100 times)
+2. Minimized Child Model
+ * decreasing computational complexity of each training (~20 times)
+3. Less stochastic augmentation search space design
+ * decreasing number of iterations needed
+
+For achieving the second goal, user interface is designed in a way that it gives user broad configuration possibilities and model selections (e.g. selecting the child model or inputting a self-designed child model).
+
+## Importance
+### Practical importance
+DeepAugment makes optimization of data augmentation scalable, and thus enables users to optimize augmentation policies without needing massive computational resources.
+As an estimate of its computational cost, it takes **4.2 hours** (500 iterations) on CIFAR-10 dataset which costs around **$13** using AWS p3.x2large instance.
+### Academic importance
+To our knowledge, DeepAugment is the first method which utilizes Bayesian Optimization for the problem of data augmentation hyperparameter optimization.
+
+## How it works
+
+Three major components of DeepAugment are controller, augmenter, and child model. Overall workflow is that controller samples new augmentation policies, augmenter transforms images by the new policy, and child model is trained from scratch by augmented images. Then, a reward is calculated from child model's training history. This reward is returned back to the controller, and it updates its surrogate model with this reward and associated augmentation policy. Then, controller samples new policies again and same steps repeats. This process cycles until user-determined maximum number of iterations reached.
+
+Controller can be set for using either Bayesian Optimization (default) or Random Search. If set to Bayesian Optimization, samples new policies by a Random Forest Estimator and Expected Improvement acquisition function.
+
+ +
+### Why Bayesian Optimization?
+
+In hyperparameter optimization, main choices are random search, grid search, bayesian optimization (BO), and reinforcement learning (RL) (in the order of method complexity). Google's [AutoAugment](https://arxiv.org/abs/1805.09501) uses RL for data augmentation hyperparameter tuning, but it takes 15,000 iterations to learn policies (which means training the child CNN model 15,000 times). Thus, it requires massive computational resources. Bayesian Optimization on the other hand learns good polices in 100-300 iterations, making it +40X faster. Additionally, it is better than grid search and random search in terms of accuracy, cost, and computation time in hyperparameter tuning([ref](https://mlconf.com/lets-talk-bayesian-optimization/)) (we can think optimization of augmentation policies as a hyperparameter tuning problem where hyperparameters are concerning with augmentations instead of the deep learning architecture). This result is not surprising since despite Grid Search or Random Search BO selects new hyperparameter as informed with previous results for tried hyperparameters.
+
+
+
+### Why Bayesian Optimization?
+
+In hyperparameter optimization, main choices are random search, grid search, bayesian optimization (BO), and reinforcement learning (RL) (in the order of method complexity). Google's [AutoAugment](https://arxiv.org/abs/1805.09501) uses RL for data augmentation hyperparameter tuning, but it takes 15,000 iterations to learn policies (which means training the child CNN model 15,000 times). Thus, it requires massive computational resources. Bayesian Optimization on the other hand learns good polices in 100-300 iterations, making it +40X faster. Additionally, it is better than grid search and random search in terms of accuracy, cost, and computation time in hyperparameter tuning([ref](https://mlconf.com/lets-talk-bayesian-optimization/)) (we can think optimization of augmentation policies as a hyperparameter tuning problem where hyperparameters are concerning with augmentations instead of the deep learning architecture). This result is not surprising since despite Grid Search or Random Search BO selects new hyperparameter as informed with previous results for tried hyperparameters.
+
+ +
+### How does Bayesian Optimization work?
+
+Aim of Bayesian Optimization (BO) is finding **set of parameters** which maximize the value of an **objective function**. It builds a surrogate model for predicting value of objective function for unexplored parameters. Working cycle of BO can be summarized as:
+1. Build a surrogate model of the objective function
+2. Find parameters that perform best on the surrogate (or pick random hyperparameters)
+3. Execute objective function with these parameters
+4. Update the surrogate model with these parameters and result (value) of objective function
+5. Repeat steps 2-4 until maximum number of iterations reached
+
+For more detailed explanation, read [this blogpost](https://towardsdatascience.com/a-conceptual-explanation-of-bayesian-model-based-hyperparameter-optimization-for-machine-learning-b8172278050f) explaining BO in high-level, or take a glance at [this review paper](https://ieeexplore.ieee.org/document/7352306)
+
+### Augmentation policy
+
+A policy describes the augmentation will be applied on a dataset. Each policy consists variables for two augmentation types, their magnitude and the portion of the data to be augmented. An example policy is as following:
+
+
+
+### How does Bayesian Optimization work?
+
+Aim of Bayesian Optimization (BO) is finding **set of parameters** which maximize the value of an **objective function**. It builds a surrogate model for predicting value of objective function for unexplored parameters. Working cycle of BO can be summarized as:
+1. Build a surrogate model of the objective function
+2. Find parameters that perform best on the surrogate (or pick random hyperparameters)
+3. Execute objective function with these parameters
+4. Update the surrogate model with these parameters and result (value) of objective function
+5. Repeat steps 2-4 until maximum number of iterations reached
+
+For more detailed explanation, read [this blogpost](https://towardsdatascience.com/a-conceptual-explanation-of-bayesian-model-based-hyperparameter-optimization-for-machine-learning-b8172278050f) explaining BO in high-level, or take a glance at [this review paper](https://ieeexplore.ieee.org/document/7352306)
+
+### Augmentation policy
+
+A policy describes the augmentation will be applied on a dataset. Each policy consists variables for two augmentation types, their magnitude and the portion of the data to be augmented. An example policy is as following:
+
+ +
+There are currently 20 types of augmentation techniques (above, right) that each aug. type variable can take. All techniques are (this list might grow in later versions):
+```Python
+AUG_TYPES = [ "crop", "gaussian-blur", "rotate", "shear", "translate-x", "translate-y", "sharpen", "emboss", "additive-gaussian-noise", "dropout", "coarse-dropout", "gamma-contrast", "brighten", "invert", "fog", "clouds", "add-to-hue-and-saturation", "coarse-salt-pepper", "horizontal-flip", "vertical-flip"]
+```
+### Child model
+[source](https://github.com/barisozmen/deepaugment/blob/master/deepaugment/childcnn.py#L232-L269)
+
+Child model is trained over and over from scratch during the optimization process. Its number of training depends on the number of iterations chosen by the user, which is expected to be around 100-300 for obtaining good results. Child model is therefore the computational bottleneck of the algorithm. With the current design, training time is ~30 seconds for 32x32 images on AWS instance p3.x2large using V100 GPU (112 TensorFLOPS). It has 1,250,858 trainable parameters for 32x32 images. Below is the diagram of child model:
+
+
+There are currently 20 types of augmentation techniques (above, right) that each aug. type variable can take. All techniques are (this list might grow in later versions):
+```Python
+AUG_TYPES = [ "crop", "gaussian-blur", "rotate", "shear", "translate-x", "translate-y", "sharpen", "emboss", "additive-gaussian-noise", "dropout", "coarse-dropout", "gamma-contrast", "brighten", "invert", "fog", "clouds", "add-to-hue-and-saturation", "coarse-salt-pepper", "horizontal-flip", "vertical-flip"]
+```
+### Child model
+[source](https://github.com/barisozmen/deepaugment/blob/master/deepaugment/childcnn.py#L232-L269)
+
+Child model is trained over and over from scratch during the optimization process. Its number of training depends on the number of iterations chosen by the user, which is expected to be around 100-300 for obtaining good results. Child model is therefore the computational bottleneck of the algorithm. With the current design, training time is ~30 seconds for 32x32 images on AWS instance p3.x2large using V100 GPU (112 TensorFLOPS). It has 1,250,858 trainable parameters for 32x32 images. Below is the diagram of child model:
+ +
+#### Other choices for child CNN model
+Standard Child model is a basic CNN where its diagram and details given above. However, you are not limited with that model. You can use your own keras model by assigning it into config dictionary as:
+```Python
+my_config = {"model": my_keras_model_object}
+deepaug = DeepAugment(my_images, my_labels, my_config)
+```
+Or use an implemented small model, such as WideResNet-40-2 (while it is bigger than Basic CNN):
+```Python
+my_config = {"model": "wrn_40_2"} # depth(40) and wideness-factor(2) can be changed. e.g. wrn_20_4
+```
+Or use a big model (not recommended unless you have massive computational resources):
+```Python
+my_config = {"model": "InceptionV3"}
+```
+```Python
+my_config = {"model": "MobileNetV2"}
+```
+
+### Reward function
+[source](https://github.com/barisozmen/deepaugment/blob/master/deepaugment/objective.py#L69-L89)
+
+Reward function is calculated as mean of K highest validation accuracies of the child model which is not smaller than corresponding training accuracy by 0.05. K can be determined by the user by updating `opt_last_n_epochs` key in config as argument to `DeepAugment()` class (K is 3 by default).
+
+## Configuration options
+
+DeepAugment can be given a config dictionary during initialization. It is expected to have following keys:
+
+* **model**: child model type. Options: "basiccnn", "inceptionv3", "mobilenetv2", "wrn_
+
+#### Other choices for child CNN model
+Standard Child model is a basic CNN where its diagram and details given above. However, you are not limited with that model. You can use your own keras model by assigning it into config dictionary as:
+```Python
+my_config = {"model": my_keras_model_object}
+deepaug = DeepAugment(my_images, my_labels, my_config)
+```
+Or use an implemented small model, such as WideResNet-40-2 (while it is bigger than Basic CNN):
+```Python
+my_config = {"model": "wrn_40_2"} # depth(40) and wideness-factor(2) can be changed. e.g. wrn_20_4
+```
+Or use a big model (not recommended unless you have massive computational resources):
+```Python
+my_config = {"model": "InceptionV3"}
+```
+```Python
+my_config = {"model": "MobileNetV2"}
+```
+
+### Reward function
+[source](https://github.com/barisozmen/deepaugment/blob/master/deepaugment/objective.py#L69-L89)
+
+Reward function is calculated as mean of K highest validation accuracies of the child model which is not smaller than corresponding training accuracy by 0.05. K can be determined by the user by updating `opt_last_n_epochs` key in config as argument to `DeepAugment()` class (K is 3 by default).
+
+## Configuration options
+
+DeepAugment can be given a config dictionary during initialization. It is expected to have following keys:
+
+* **model**: child model type. Options: "basiccnn", "inceptionv3", "mobilenetv2", "wrn_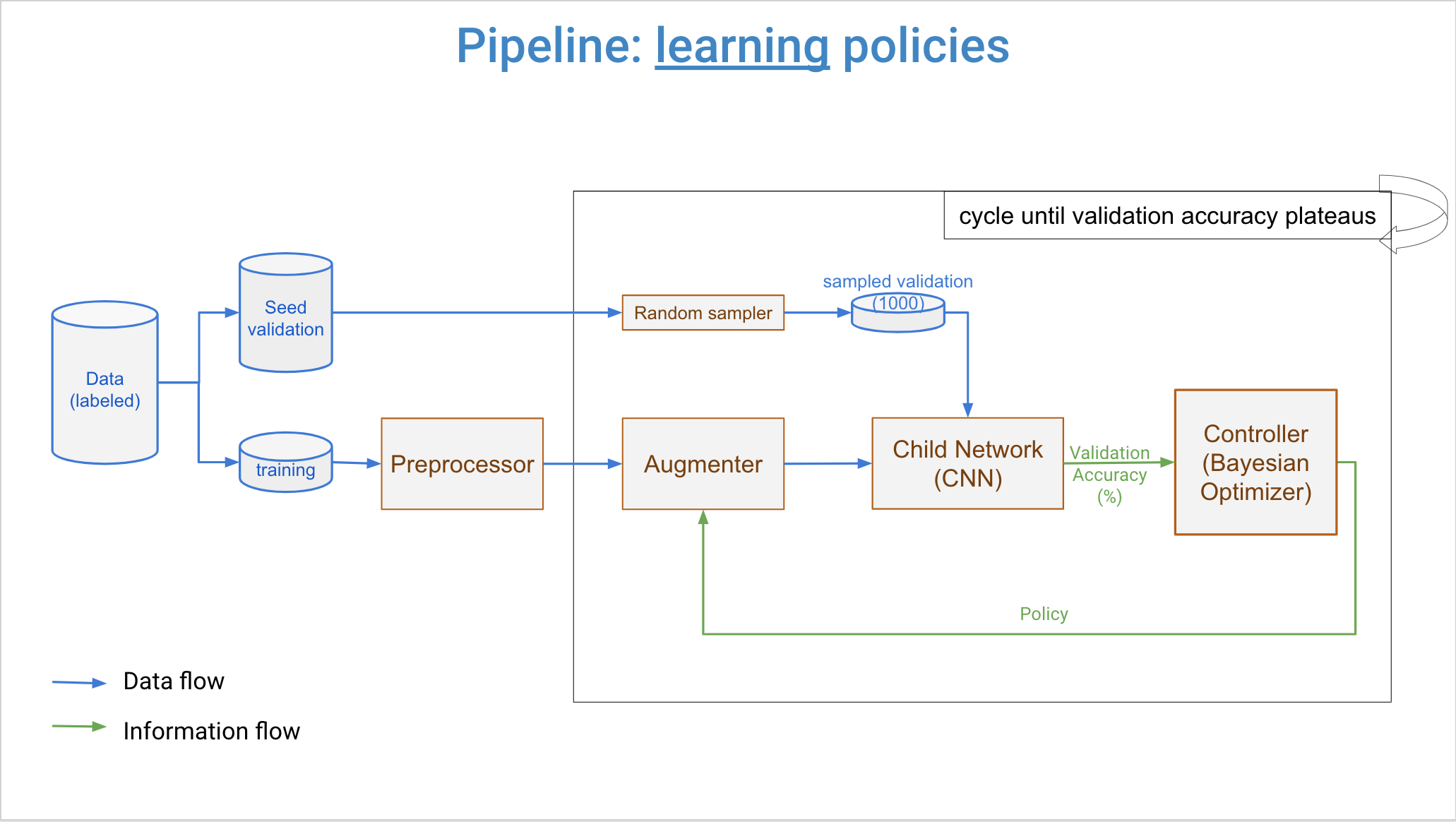 +
+ +
+## Class diagram
+Created by [pyreverse](https://www.logilab.org/blogentry/6883)
+
+
+## Package diagram
+Created by [pyreverse](https://www.logilab.org/blogentry/6883)
+
+
+## Class diagram
+Created by [pyreverse](https://www.logilab.org/blogentry/6883)
+
+
+## Package diagram
+Created by [pyreverse](https://www.logilab.org/blogentry/6883)
+ +
+## References
+[1] Cubuk et al., 2018. AutoAugment: Learning Augmentation Policies from Data
+([arxiv](https://arxiv.org/abs/1805.09501))
+
+[2] Zoph et al., 2016. Neural Architecture Search with Reinforcement Learning
+([arxiv](https://arxiv.org/abs/1611.01578))
+
+[3] Shahriari et al., 2016. A review of Bayesian Optimization
+([ieee](https://ieeexplore.ieee.org/document/7352306))
+
+[4] Dewancker et al. Bayesian Optimization Primer ([white-paper](https://app.sigopt.com/static/pdf/SigOpt_Bayesian_Optimization_Primer.pdf))
+
+[5] DeVries, Taylor 2017. Improved Regularization of CNN's with Cutout
+([arxiv](https://arxiv.org/abs/1708.04552))
+
+Blogs:
+- A conceptual explanation of Bayesian Optimization ([towardsdatascience](https://towardsdatascience.com/a-conceptual-explanation-of-bayesian-model-based-hyperparameter-optimization-for-machine-learning-b8172278050f))
+- Comparison experiment: Bayesian Opt. vs Grid Search vs Random Search ([mlconf](https://mlconf.com/lets-talk-bayesian-optimization/))
+
+Libraries:
+- [scikit-optimize](https://scikit-optimize.github.io/)
+- [imgaug](https://github.com/aleju/imgaug)
+- [AutoAugment-unofficial](https://github.com/barisozmen/autoaugment-unofficial)
+- [Automold](https://github.com/UjjwalSaxena/Automold--Road-Augmentation-Library) (Self-driving car image-augmentation library)
--------
-
+
+## References
+[1] Cubuk et al., 2018. AutoAugment: Learning Augmentation Policies from Data
+([arxiv](https://arxiv.org/abs/1805.09501))
+
+[2] Zoph et al., 2016. Neural Architecture Search with Reinforcement Learning
+([arxiv](https://arxiv.org/abs/1611.01578))
+
+[3] Shahriari et al., 2016. A review of Bayesian Optimization
+([ieee](https://ieeexplore.ieee.org/document/7352306))
+
+[4] Dewancker et al. Bayesian Optimization Primer ([white-paper](https://app.sigopt.com/static/pdf/SigOpt_Bayesian_Optimization_Primer.pdf))
+
+[5] DeVries, Taylor 2017. Improved Regularization of CNN's with Cutout
+([arxiv](https://arxiv.org/abs/1708.04552))
+
+Blogs:
+- A conceptual explanation of Bayesian Optimization ([towardsdatascience](https://towardsdatascience.com/a-conceptual-explanation-of-bayesian-model-based-hyperparameter-optimization-for-machine-learning-b8172278050f))
+- Comparison experiment: Bayesian Opt. vs Grid Search vs Random Search ([mlconf](https://mlconf.com/lets-talk-bayesian-optimization/))
+
+Libraries:
+- [scikit-optimize](https://scikit-optimize.github.io/)
+- [imgaug](https://github.com/aleju/imgaug)
+- [AutoAugment-unofficial](https://github.com/barisozmen/autoaugment-unofficial)
+- [Automold](https://github.com/UjjwalSaxena/Automold--Road-Augmentation-Library) (Self-driving car image-augmentation library)
--------
-